Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Tagami, Susumu; Shimizu, Osamu; Sano, Naruto; Imaizumi, Akiko; Tobita, Hiroshi; Nagasaki, Yosuke; Kurobane, Shiro
JAEA-Review 2010-079, 90 Pages, 2011/03
The Back-end Cycle Key Elements Research Facility (BECKY) was installed in the Nuclear Fuel Cycle Safety Engineering Research Facility (NUCEF) in 1994 for the safety and basic studies of the nuclear fuel cycle and radioactive waste. BECKY consists of three alpha/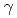 concrete cells, three alpha/
concrete cells, three alpha/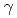 steel cells, thirty glove boxes and twenty hoods. Operation, maintenance and management are in execution based on the operational safety programs. And we also execute application about amendment of the permission and approval for the purpose of support R&D. This report describes the operation, maintenance and management from April 1, 2005 to March 31, 2010 for the purpose of keeping the performance of BECKY.
steel cells, thirty glove boxes and twenty hoods. Operation, maintenance and management are in execution based on the operational safety programs. And we also execute application about amendment of the permission and approval for the purpose of support R&D. This report describes the operation, maintenance and management from April 1, 2005 to March 31, 2010 for the purpose of keeping the performance of BECKY.
Isayama, Akihiko; Sakakibara, Satoru*; Furukawa, Masaru*; Matsunaga, Go; Yamazaki, Kozo*; Watanabe, Kiyomasa*; Idomura, Yasuhiro; Sakamoto, Yoshiteru; Tanaka, Kenji*; Tamura, Naoki*; et al.
Purazuma, Kaku Yugo Gakkai-Shi, 86(6), p.374 - 377, 2010/06
no abstracts in English
Osakabe, Masaki*; Shinohara, Koji; Toi, Kazuo*; Todo, Yasushi*; Hamamatsu, Kiyotaka; Murakami, Sadayoshi*; Yamamoto, Satoshi*; Idomura, Yasuhiro; Sakamoto, Yoshiteru; Tanaka, Kenji*; et al.
Purazuma, Kaku Yugo Gakkai-Shi, 85(12), p.839 - 842, 2009/12
no abstracts in English
Idomura, Yasuhiro; Yoshida, Maiko; Yagi, Masatoshi*; Tanaka, Kenji*; Hayashi, Nobuhiko; Sakamoto, Yoshiteru; Tamura, Naoki*; Oyama, Naoyuki; Urano, Hajime; Aiba, Nobuyuki; et al.
Purazuma, Kaku Yugo Gakkai-Shi, 84(12), p.952 - 955, 2008/12
no abstracts in English
Yamasaki, Chisato*; Murakami, Katsuhiko*; Fujii, Yasuyuki*; Sato, Yoshiharu*; Harada, Erimi*; Takeda, Junichi*; Taniya, Takayuki*; Sakate, Ryuichi*; Kikugawa, Shingo*; Shimada, Makoto*; et al.
Nucleic Acids Research, 36(Database), p.D793 - D799, 2008/01
Times Cited Count:52 Percentile:71.15(Biochemistry & Molecular Biology)Here we report the new features and improvements in our latest release of the H-Invitational Database, a comprehensive annotation resource for human genes and transcripts. H-InvDB, originally developed as an integrated database of the human transcriptome based on extensive annotation of large sets of fulllength cDNA (FLcDNA) clones, now provides annotation for 120 558 human mRNAs extracted from the International Nucleotide Sequence Databases (INSD), in addition to 54 978 human FLcDNAs, in the latest release H-InvDB. We mapped those human transcripts onto the human genome sequences (NCBI build 36.1) and determined 34 699 human gene clusters, which could define 34 057 protein-coding and 642 non-protein-coding loci; 858 transcribed loci overlapped with predicted pseudogenes.