Protein conformation and its dynamics in solution by molecular dynamics simulation for small-angle neutron scattering data analysis
Putra, E. G. R.*; Kono, Hidetoshi; Tokuhisa, Atsushi*; Bahrum, E. S.*; Patriati, A.*
Molecular dynamics (MD) simulation of hen egg white lysozyme protein (4LZT, with and without disulphide bonds) and Ca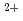 -binding protein calmodulin (1PRW, bent conformation and 1EXR, extended conformation) in solution have been explored using computer simulation. The dynamics of the proteins in aqueous phase at room temperature was produced for 5 nanosecond (ns) time-scale calculation. The trajectories generated from MD simulation were computed to produce the pair distribution function (PDF), the distribution of the neutron scattering length density as a function of atomic distances of a single molecule protein. Conclusively an inverse Fourier transform was applied to transform a pair-distribution function data into a theoretical small-angle neutron scattering (SANS) intensity against scattering vector (q) which can be experimentally evaluated. Several information such as the radius of gyration (Rg) and the shape of protein in solution can be obtained at every single time of simulation. In this study, the time-dependent structural changes in the extended conformation of Ca
-binding protein calmodulin (1PRW, bent conformation and 1EXR, extended conformation) in solution have been explored using computer simulation. The dynamics of the proteins in aqueous phase at room temperature was produced for 5 nanosecond (ns) time-scale calculation. The trajectories generated from MD simulation were computed to produce the pair distribution function (PDF), the distribution of the neutron scattering length density as a function of atomic distances of a single molecule protein. Conclusively an inverse Fourier transform was applied to transform a pair-distribution function data into a theoretical small-angle neutron scattering (SANS) intensity against scattering vector (q) which can be experimentally evaluated. Several information such as the radius of gyration (Rg) and the shape of protein in solution can be obtained at every single time of simulation. In this study, the time-dependent structural changes in the extended conformation of Ca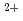 -binding protein calmodulin every 100 picosecond (ps) presented in through the pair-distribution function and SANS scattering profiles.
-binding protein calmodulin every 100 picosecond (ps) presented in through the pair-distribution function and SANS scattering profiles.