Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Tamada, Taro; Adachi, Motoyasu; Kurihara, Kazuo; Kuroki, Ryota
Nihon Kessho Gakkai-Shi, 55(1), p.47 - 51, 2013/02
Neutron crystallography enables us to identify the accurate hydrogen positions in proteins, which play important roles in many chemical reactions in living system. Here we show our results of neutron structure determination of enzymes in complex with its inhibitors which corresponds to transition state analogues. Neutron structure analysis elucidated the detail catalytic reaction of each enzyme by direct observation of hydrogen atoms. Furthermore we would like to introduce a new neutron beam line for neutron structural biology planned at MLF in J-PARC.
 HB8
HB8Okazaki, Nobuo; Adachi, Motoyasu; Tamada, Taro; Kurihara, Kazuo; Oga, Takushi*; Kamiya, Nobuo*; Kuramitsu, Seiki*; Kuroki, Ryota
Acta Crystallographica Section F, 68(1), p.49 - 52, 2012/01
Times Cited Count:2 Percentile:32.79(Biochemical Research Methods)Kuroki, Ryota; Okazaki, Nobuo; Adachi, Motoyasu; Ohara, Takashi; Kurihara, Kazuo; Tamada, Taro
Acta Crystallographica Section D, 66(11), p.1126 - 1130, 2010/11
Times Cited Count:2 Percentile:30.11(Biochemical Research Methods)It is generally known that enzymes represent important drug-target proteins. Elucidation of the catalytic function and the molecular-recognition mechanisms of enzymes provides important information for structure-based drug design. Neutron crystallography provides accurate information on the locations of H atoms that are essential in enzymatic function and molecular recognition. Recent examples are described of the structure determination of the drug-target proteins human immunodeficiency virus protease and porcine pancreatic elastase in complex with transition-state analogue inhibitors using the neutron diffractometers for biological crystallography (BIX-3 and BIX-4) installed at the JRR-3 research reactor.
Kuroki, Ryota; Tamada, Taro; Kurihara, Kazuo; Ohara, Takashi; Adachi, Motoyasu
Yakugaku Zasshi, 130(5), p.657 - 664, 2010/05
Times Cited Count:0 Percentile:0.02(Pharmacology & Pharmacy)Crystallography enables us to obtain accurate atomic positions within proteins. High resolution X-ray crystallography provides information for most of the atoms comprising a protein, with the exception of hydrogens. Neutron diffraction data can provide information of the location of hydrogen atoms to the structural information determined by X-ray crystallography. Here, we show the recent result of the structural determination of drug-target proteins, porcine pancreatic elastase and human immuno-deficiency virus type-1 protease by both X-ray and neutron diffraction. The structure of porcine pancreatic elastase with its potent inhibitor was determined at room temperature to 1.2  resolution by X-ray diffraction and 1.65
resolution by X-ray diffraction and 1.65  resolution by neutron diffraction. The structure of HIV-PR with its potent inhibitor was also determined to 1.4
resolution by neutron diffraction. The structure of HIV-PR with its potent inhibitor was also determined to 1.4  resolution by X-ray diffraction and 1.9
resolution by X-ray diffraction and 1.9  resolution by neutron diffraction. Ultra-high resolution structures of both proteins (0.94
resolution by neutron diffraction. Ultra-high resolution structures of both proteins (0.94  and 0.93
and 0.93  , respectively) were also determined by X-ray diffraction at 100 K. The ionization state and the location of hydrogen atoms of the catalytic residue in these enzymes were determined by neutron diffraction. Furthermore, collaborative use of both X-ray and neutron to identify the location of ambiguous hydrogen atoms will be shown.
, respectively) were also determined by X-ray diffraction at 100 K. The ionization state and the location of hydrogen atoms of the catalytic residue in these enzymes were determined by neutron diffraction. Furthermore, collaborative use of both X-ray and neutron to identify the location of ambiguous hydrogen atoms will be shown.
Tamada, Taro; Kinoshita, Takayoshi*; Kurihara, Kazuo; Adachi, Motoyasu; Ohara, Takashi; Imai, Keisuke*; Kuroki, Ryota; Tada, Toshiji*
Journal of the American Chemical Society, 131(31), p.11033 - 11040, 2009/07
Times Cited Count:58 Percentile:79.01(Chemistry, Multidisciplinary)To help resolve long-standing questions regarding the catalytic activity of the serine proteases the structure of porcine pancreatic elastase has been analyzed by high-resolution neutron and X-ray crystallography. In order to mimic the tetrahedral transition intermediate a peptidic inhibitor was used. A single large crystal was used to collect room-temperature neutron data to 1.65  resolution and X-ray data to 1.20
resolution and X-ray data to 1.20  resolution. Another crystal provided a low-temperature X-ray data set to 0.94
resolution. Another crystal provided a low-temperature X-ray data set to 0.94  resolution. The neutron data are to higher resolution than previously reported for a serine protease and the X-ray data are comparable with other studies. The neutron and X-ray data show that the hydrogen bond between His57 and Asp102 (chymotrypsin numbering) is 2.60
resolution. The neutron data are to higher resolution than previously reported for a serine protease and the X-ray data are comparable with other studies. The neutron and X-ray data show that the hydrogen bond between His57 and Asp102 (chymotrypsin numbering) is 2.60  in length and that the hydrogen-bonding hydrogen is 0.80-0.96
in length and that the hydrogen-bonding hydrogen is 0.80-0.96  from the histidine nitrogen. This is not consistent with a low-barrier hydrogen which is predicted to have the hydrogen midway between the donor and acceptor atom. The observed interaction between His57 and Asp102 is essentially a short but conventional hydrogen bond, sometimes described as a short ionic hydrogen bond. The neutron analysis also shows that the oxygen of the oxopropyl group of the inhibitor is present as an oxygen anion rather than a hydroxyl group, supporting the role of the "oxyanion hole" in stabilizing the tetrahedral intermediate in catalysis.
from the histidine nitrogen. This is not consistent with a low-barrier hydrogen which is predicted to have the hydrogen midway between the donor and acceptor atom. The observed interaction between His57 and Asp102 is essentially a short but conventional hydrogen bond, sometimes described as a short ionic hydrogen bond. The neutron analysis also shows that the oxygen of the oxopropyl group of the inhibitor is present as an oxygen anion rather than a hydroxyl group, supporting the role of the "oxyanion hole" in stabilizing the tetrahedral intermediate in catalysis.
Adachi, Motoyasu; Ohara, Takashi; Kurihara, Kazuo; Tamada, Taro; Honjo, Eijiro; Okazaki, Nobuo; Arai, Shigeki; Shoyama, Yoshinari; Kimura, Kaname*; Matsumura, Hiroyoshi*; et al.
Proceedings of the National Academy of Sciences of the United States of America, 106(12), p.4641 - 4646, 2009/03
Times Cited Count:111 Percentile:90.72(Multidisciplinary Sciences)To further understand the catalytic mechanism and inhibitor recognition of HIV-1 protease, we need to determine the locations of key hydrogen atoms in the catalytic aspartates Asp25 and Asp125. The structure of HIV-1 protease in complex with transition-state analog KNI-272 was determined by combined neutron crystallography at 1.9  resolution and X-ray crystallography at 1.4
resolution and X-ray crystallography at 1.4  resolution. The resulting structural data shows that the catalytic residue Asp25 is protonated and that Asp125 is deprotonated. The proton on Asp25 makes a hydrogen bond with the carbonyl group of the allophenylnorstatine group in KNI-272. The deprotonated Asp125 bonds to the hydroxyl proton of Apns. The results provide direct experimental evidence for proposed aspects of the catalytic mechanism of HIV-1 protease; and can therefore contribute substantially to the development of specific inhibitors for therapeutic application.
resolution. The resulting structural data shows that the catalytic residue Asp25 is protonated and that Asp125 is deprotonated. The proton on Asp25 makes a hydrogen bond with the carbonyl group of the allophenylnorstatine group in KNI-272. The deprotonated Asp125 bonds to the hydroxyl proton of Apns. The results provide direct experimental evidence for proposed aspects of the catalytic mechanism of HIV-1 protease; and can therefore contribute substantially to the development of specific inhibitors for therapeutic application.
Adachi, Jun*; Kurosaki, Ken*; Uno, Masayoshi*; Yamanaka, Shinsuke*; Takano, Masahide; Akabori, Mitsuo; Minato, Kazuo
Journal of Nuclear Materials, 384(1), p.6 - 11, 2009/01
Times Cited Count:9 Percentile:53.3(Materials Science, Multidisciplinary)The indentation hardness, Vickers hardness, fracture toughness, and Young's modulus of polycrystalline uranium monontride (UN) at sub-microscale and macroscale were evaluated by an indentation test, Vickers hardness test, and the ultrasonic pulse echo method. The Young's modulus and Vickers hardness were in good agreement with the literature values. The fracture toughness of UN was about three times that of UO . In addition, we revealed the indentation size effect on the indentation hardness of UN.
. In addition, we revealed the indentation size effect on the indentation hardness of UN.
Takamura, Shuichi*; Kado, Shinichiro*; Fujii, Takashi*; Fujiyama, Hiroshi*; Takabe, Hideaki*; Adachi, Kazuo*; Morimiya, Osamu*; Fujimori, Naoji*; Watanabe, Takayuki*; Hayashi, Yasuaki*; et al.
Kara Zukai, Purazuma Enerugi No Subete, P. 164, 2007/03
no abstracts in English
Hanzawa, Yukiko; Magara, Masaaki; Watanabe, Kazuo; Esaka, Fumitaka; Miyamoto, Yutaka; Yasuda, Kenichiro; Gunji, Katsubumi*; Yamamoto, Yoichi; Takahashi, Tsukasa; Sakurai, Satoshi; et al.
JAERI-Tech 2002-103, 141 Pages, 2003/02
The JAERI has established a facility with a cleanroom: the Clean Laboratory for Environmental Analysis and Research (CLEAR). This report is an overview of the design, construction and performance evaluation of the CLEAR in the initial stage of the laboratory operation in June 2001. The CLEAR is a facility to be used for ultra trace analyses of nuclear materials in environmental samples for the safeguards, for the CTBT verification and for researches on environmental sciences. The CLEAR meets double requirements of a cleanroom and for handling of nuclear materials. Much attention was paid to the construction materials of the cleanroom for trace analysis of metal elements using corrosive acids. The air conditioning and purification system, experimental equipment, utilities and safety systems are also demonstrated. The potential contamination from the completed cleanroom atmosphere during the analytical procedure was evaluated. It can be concluded that the CLEAR has provided a suitable condition for reliable analysis of ultra trace amounts of nuclear materials in environmental samples.
Idomura, Yasuhiro; Adachi, Masaaki*; Gorai, Kazuo; Suzuki, Yoshio; Wang, X.*
Purazuma, Kaku Yugo Gakkai-Shi, 79(2), p.172 - 187, 2003/02
Under the Numerical EXperiment of Tokamak (NEXT) research project, various fluid, particle, and hybrid codes have been developed. These codes require a computational environment which consists of high performance processors, high speed storage system, and high speed parallelized visualization system. In this paper, the performance of the JAERI Origin3800 system is examined from a point of view of these requests. In the performance tests, it is shown that the representative particle and fluid codes operate with 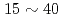 % of processing efficiency up to 512 processors. A storage area network (SAN) provides high speed parallel data transfer. A parallel visualization system enables order of magnitude faster visualization of a large scale simulation data compared with the previous graphic workstations. Accordingly, an extremely advanced simulation environment is realized on the JAERI Origin3800 system. Recently, development of a storage grid is underway in order to improve a computational environment of remote users. The storage grid is constructed by a combination of SAN and a wavelength division multiplexer (WDM). The preliminary tests show that compared with the existing data transfer methods, it enables dramatically high speed data transfer
% of processing efficiency up to 512 processors. A storage area network (SAN) provides high speed parallel data transfer. A parallel visualization system enables order of magnitude faster visualization of a large scale simulation data compared with the previous graphic workstations. Accordingly, an extremely advanced simulation environment is realized on the JAERI Origin3800 system. Recently, development of a storage grid is underway in order to improve a computational environment of remote users. The storage grid is constructed by a combination of SAN and a wavelength division multiplexer (WDM). The preliminary tests show that compared with the existing data transfer methods, it enables dramatically high speed data transfer 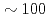 Gbps over a wide area network.
Gbps over a wide area network.
Hanzawa, Yukiko; Magara, Masaaki; Watanabe, Kazuo; Esaka, Fumitaka; Miyamoto, Yutaka; Yasuda, Kenichiro; Gunji, Katsubumi*; Sakurai, Satoshi; Takano, Seinojo*; Usuda, Shigekazu; et al.
Journal of Nuclear Science and Technology, 40(1), p.49 - 56, 2003/01
Times Cited Count:4 Percentile:31.64(Nuclear Science & Technology)The JAERI has established a cleanroom facility with cleanliness of ISO Class 5: the Clean Laboratory for Environmental Analysis and Research (CLEAR). It was designed to be used for the analysis of nuclear materials in environmental samples for the safeguards, the Comprehensive Nuclear-Test-Ban Treaty verification and research on environmental sciences. The CLEAR facility was designed to meet double conflicting requirements of a cleanroom and for handling of nuclear materials according to Japanese regulations, i.e., to avoid contamination from outside and to contain nuclear materials inside the facility. This facility has been intended to be used for wet chemical treatment, instrumental analysis and particle handling. A fume-hood to provide a clean work surface for handling of nuclear materials was specially designed. The performance of the cleanroom and analytical background in the laboratory are discussed. It can be concluded that the CLEAR facility enables analysis of ultra trace amounts of nuclear materials at the sub-picogram level in environmental samples.
Sakurai, Satoshi; Hanzawa, Yukiko; Magara, Masaaki; Usuda, Shigekazu; Watanabe, Kazuo; Adachi, Takeo
Kuki Seijo, 39(6), p.404 - 410, 2002/03
no abstracts in English
Esaka, Fumitaka; Magara, Masaaki; Hanzawa, Yukiko; Sakurai, Satoshi; Taguchi, Takuji; Takai, Konomi; Sakakibara, Takaaki; Kurosawa, Setsumi; Takahashi, Masato; Yasuda, Kenichiro; et al.
Dai-22-Kai Kaku Busshitsu Kanri Gakkai Nihon Shibu Nenji Taikai Rombunshu, 8 Pages, 2001/11
no abstracts in English
Usuda, Shigekazu; Watanabe, Kazuo; Sakurai, Satoshi; Magara, Masaaki; Hanzawa, Yukiko; Esaka, Fumitaka; Miyamoto, Yutaka; Yasuda, Kenichiro; Saito, Yoko; Gunji, Katsubumi*; et al.
KEK Proceedings 2001-14, p.88 - 92, 2001/06
no abstracts in English
Esaka, Fumitaka; Zheng, W.*; Watanabe, Kazuo; Magara, Masaaki; Hanzawa, Yukiko; Usuda, Shigekazu; Adachi, Takeo
Advances in Mass Spectrometry, 15, p.973 - 974, 2001/00
no abstracts in English
Adachi, Takeo; Usuda, Shigekazu; Watanabe, Kazuo; Sakurai, Satoshi; Magara, Masaaki; Hanzawa, Yukiko; Esaka, Fumitaka; Yasuda, Kenichiro; Saito, Yoko; Takahashi, Masato; et al.
IAEA-SM-367/10/02 (CD-ROM), 8 Pages, 2001/00
no abstracts in English
Magara, Masaaki; Hanzawa, Yukiko; Esaka, Fumitaka; Miyamoto, Yutaka; Yasuda, Kenichiro; Watanabe, Kazuo; Usuda, Shigekazu; Nishimura, Hideo; Adachi, Takeo
Applied Radiation and Isotopes, 53(1-2), p.87 - 90, 2000/07
Times Cited Count:28 Percentile:84.32(Chemistry, Inorganic & Nuclear)no abstracts in English
Nishimura, Hideo; Magara, Masaaki; Hanzawa, Yukiko; Esaka, Fumitaka; Takahashi, Tsukasa; Gunji, Katsubumi; Miyamoto, Yutaka; Yasuda, Kenichiro; Tsuruta, Yasuhiro; Tsuda, Shinji; et al.
Heisei 11-Nendo Hosho Sochi Semina Koenroku Tekisuto, p.95 - 107, 2000/01
no abstracts in English
Esaka, Fumitaka; Watanabe, Kazuo; Magara, Masaaki; Hanzawa, Yukiko; Usuda, Shigekazu; Gunji, Katsubumi; Nishimura, Hideo; Adachi, Takeo
Proceedings of 12th International Conference on Secondary Ion Mass Spectrometry (SIMS 12), p.977 - 980, 2000/00
no abstracts in English
Usuda, Shigekazu; Adachi, Takeo; Watanabe, Kazuo; Magara, Masaaki; Hanzawa, Yukiko; Esaka, Fumitaka; Miyamoto, Yutaka; Yasuda, Kenichiro; Gunji, Hideho; Tsuruta, Yasuhiro; et al.
Proceedings of Seminar on Strengthening of Safeguards: Integrating the New and the Old, p.477 - 481, 2000/00
no abstracts in English