Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
天野 由記; Sachdeva, R.*; Gittins, D.*; Anantharaman, K.*; Lei, S.*; Valentin-Alvarado, L. E.*; Diamond, S.*; 別部 光里*; 岩月 輝希; 望月 陽人; et al.
Environmental Microbiome (Internet), 19, p.105_1 - 105_17, 2024/12
被引用回数:0 パーセンタイル:0.00(Genetics & Heredity)Underground research laboratories (URLs) provide a window on the deep biosphere and enable investigation of potential microbial impacts on nuclear waste, CO and H
and H stored in the subsurface. We carried out the first multi-year study of groundwater microbiomes sampled from defined intervals between 140 and 400 m below the surface of the Horonobe and Mizunami URLs, Japan. The Horonobe and Mizunami microbiomes are dissimilar, likely because the Mizunami URL is hosted in granitic rock and the Horonobe URL in sedimentary rock. Despite this, hydrogen metabolism, rubisco-based CO
stored in the subsurface. We carried out the first multi-year study of groundwater microbiomes sampled from defined intervals between 140 and 400 m below the surface of the Horonobe and Mizunami URLs, Japan. The Horonobe and Mizunami microbiomes are dissimilar, likely because the Mizunami URL is hosted in granitic rock and the Horonobe URL in sedimentary rock. Despite this, hydrogen metabolism, rubisco-based CO fixation, reduction of nitrogen compounds and sulfate reduction are well represented functions in microbiomes from both URLs, although methane metabolism is more prevalent at the organic- and CO
fixation, reduction of nitrogen compounds and sulfate reduction are well represented functions in microbiomes from both URLs, although methane metabolism is more prevalent at the organic- and CO -rich Horonobe URL. We detected near-identical genotypes for approximately one third of all genomically defined organisms at multiple depths within the Horonobe URL. This cannot be explained by inactivity, as in situ growth was detected for some bacteria, albeit at slow rates. Given the current low hydraulic conductivity and groundwater compositional heterogeneity, ongoing inter-site strain dispersal seems unlikely. Alternatively, the Horonobe URL microbiome homogeneity may be explained by higher groundwater mobility during the last glacial period. Genotypically-defined species closely related to those detected in the URLs were identified in three other subsurface environments in the USA. Thus, dispersal rates between widely separated underground sites may be fast enough relative to mutation rates to have precluded substantial divergence in species composition. Species overlaps between subsurface locations on different continents constrain expectations regarding the scale of global subsurface biodiversity. Overall, microbiome and geochemical stability over the study period has important implications for underground storage applications.
-rich Horonobe URL. We detected near-identical genotypes for approximately one third of all genomically defined organisms at multiple depths within the Horonobe URL. This cannot be explained by inactivity, as in situ growth was detected for some bacteria, albeit at slow rates. Given the current low hydraulic conductivity and groundwater compositional heterogeneity, ongoing inter-site strain dispersal seems unlikely. Alternatively, the Horonobe URL microbiome homogeneity may be explained by higher groundwater mobility during the last glacial period. Genotypically-defined species closely related to those detected in the URLs were identified in three other subsurface environments in the USA. Thus, dispersal rates between widely separated underground sites may be fast enough relative to mutation rates to have precluded substantial divergence in species composition. Species overlaps between subsurface locations on different continents constrain expectations regarding the scale of global subsurface biodiversity. Overall, microbiome and geochemical stability over the study period has important implications for underground storage applications.
Al-Shayeb, B.*; Sachdeva, R.*; Chen, L.-X.*; Ward, F.*; Munk, P.*; Devoto, A.*; Castelle, C. J.*; Olm, M. R.*; Bouma-Gregson, K.*; 天野 由記; et al.
Nature, 578(7795), p.425 - 431, 2020/02
被引用回数:291 パーセンタイル:99.46(Multidisciplinary Sciences)Phage typically have small genomes and depend on their bacterial hosts for replication. We generated metagenomic datasets from many diverse ecosystems and reconstructed hundreds of huge phage genomes, between 200 kbp and 716 kbp in length. Thirty four genomes were manually curated to completion, including the largest phage genomes yet reported. Expanded genetic repertoires include diverse and new CRISPR-Cas systems, tRNAs, tRNA synthetases, tRNA modification enzymes, initiation and elongation factors and ribosomal proteins. Phage CRISPR have the capacity to silence host transcription factors and translational genes, potentially as part of a larger interaction network that intercepts translation to redirect biosynthesis to phage-encoded functions. Some phage repurpose bacterial systems for phage-defense to eliminate competing phage. We phylogenetically define seven major clades of huge phage from human and other animal microbiomes, oceans, lakes, sediments, soils and the built environment. We conclude that large gene inventories reflect a conserved biological strategy, observed across a broad bacterial host range and resulting in the distribution of huge phage across Earth's ecosystems.
Matheus Carnevali, P. B.*; Schulz, F.*; Castelle, C. J.*; Kantor, R. S.*; Shih, P.*; Sharon, I.*; Santini, J.*; Olm, M. R.*; 天野 由記; Thomas, B. C.*; et al.
Nature Communications (Internet), 10, p.463_1 - 463_15, 2019/01
被引用回数:48 パーセンタイル:88.22(Multidisciplinary Sciences)The metabolic platform in which microbial aerobic respiration evolved is tightly linked to the origins of Cyanobacteria (Oxyphotobacteria). Melainabacteria and Sericytochromatia, close phylogenetic neighbores to Oxyphotobacteria comprise both fermentative and aerobic representatives, or clades that are capablee of both. Here, we predict the metabolisms of Margulisbacteria from two distinct environments and Saganbacteria, and compare them to genomes of organisms from the related lineages. Melainabacteria BJ4A obtained from Mizunami site are potentially able to use O and other terminal electron acceptors. The type C heme-copper oxygen reductase found in Melainabacteria BJ4A may be adapted to low O
and other terminal electron acceptors. The type C heme-copper oxygen reductase found in Melainabacteria BJ4A may be adapted to low O levels, as expected for microaerophilic or anoxic environments such as the subsurface. Notably, Melainabacteria BJ4A seems to have a branched electron transport chain, with one branch leading to a cytochrome d ubiquinol oxidoreductase and the other one leading to the type C heme-copper oxygen reductase. Both these enzymes have high affinity for O
levels, as expected for microaerophilic or anoxic environments such as the subsurface. Notably, Melainabacteria BJ4A seems to have a branched electron transport chain, with one branch leading to a cytochrome d ubiquinol oxidoreductase and the other one leading to the type C heme-copper oxygen reductase. Both these enzymes have high affinity for O , thus are adapted to low O
, thus are adapted to low O levels. These contemporary lineages have representatives with fermentative H
levels. These contemporary lineages have representatives with fermentative H -based metabolism, lineages capable of aerobic or anaerobic respiration, and lineages with both. Our findings support the idea that the ancestor of these lineages was an anaerobe in which fermentation and H
-based metabolism, lineages capable of aerobic or anaerobic respiration, and lineages with both. Our findings support the idea that the ancestor of these lineages was an anaerobe in which fermentation and H metabolism were central metabolic features.
metabolism were central metabolic features.
 and metal transformations associated with novel bacteria and archaea in deep terrestrial subsurface sediments
and metal transformations associated with novel bacteria and archaea in deep terrestrial subsurface sedimentsHernsdorf, A. W.*; 天野 由記; 宮川 和也; 伊勢 孝太郎; 鈴木 庸平*; Anantharaman, K.*; Probst, A. J.*; Burstein, D.*; Thomas, B. C.*; Banfield, J. F.*
ISME Journal, 11, p.1915 - 1929, 2017/03
被引用回数:106 パーセンタイル:96.25(Ecology)地層処分システムにおける微生物影響の可能性を評価するために、北海道の幌延深地層研究センター地下施設を利用して、堆積岩地下の生態系における微生物群集構造と代謝機能について調査を行った。全体として、微生物生態系は多様な系統群からなる微生物種で構成されており、その多くはこれまで培養されていない生物門に属していることが示された。大部分の微生物種は、酸化型[NiFe]ヒドロゲナーゼあるいはフェレドキシンをベースとする代謝経路を可能にする電子分岐型[FeFe]ヒドロゲナーゼを介して水素代謝をおこなうことが明らかになった。水素代謝と関連して、多くの微生物が炭素,窒素,鉄および硫黄を代謝することが推定された。特に、ANME-2dというメタン酸化を行う古細菌として知られている未培養微生物が、鉄関連の代謝反応を行う可能性が示唆された。得られた結果から、幌延堆積岩環境における微生物群集の生態学的概念モデルを推定した。
Hug, L. A.*; Baker, B. J.*; Anantharaman, K.*; Brown, C. T.*; Probst, A. J.*; Castelle, C. J.*; Butterfield, C. N.*; Hernsdorf, A. W.*; 天野 由記; 伊勢 孝太郎; et al.
Nature Microbiology (Internet), 1(5), p.16048_1 - 16048_6, 2016/05
被引用回数:1364 パーセンタイル:99.96(Microbiology)生命の系統樹は生物学において最も重要な中心テーマの一つである。遺伝子調査によると、莫大な数のブランチの存在が示唆されているが、フルスケールに近い系統樹でさえわかりにくいのが現状である。本研究では、これまでに報告されてきた配列情報に加えて、新たに取得した未培養生物のゲノム情報を用いて、バクテリア,アーキア,真核生物を含む系統樹を更新した。系統樹の描写は、全体的な概容とそれぞれの主要な系統における多様性のスナップショットの両方について行った。その結果、バクテリアの多様化の優勢性が示され、培養されていない生物種の重要性とともに主要な放射構造においてそれらの生物種の重要な進化が集中している現象が強調された。
 fixation by abundant Nitrospirae in the deep subsurface
fixation by abundant Nitrospirae in the deep subsurface天野 由記; Anantharaman, K.*; Tomas, B. C.*; Olm, M.*; Burstein, D.*; Castelle, C. J.*; 別部 光里*; 宮川 和也; 岩月 輝希; 鈴木 庸平*; et al.
no journal, ,
The bacterial phylum Nitrospirae is phylogenetically diverse. There are relatively few isolated representatives available for laboratory study and the physiology, functions and distributions of these bacteria across environments remain largely unknown. To understand the ecological role of Nitrospirae in the deep subsurface, we analyzed metagenomically-derived near complete genomes from groundwaters associated with granite and sedimentary rock where Nitrospirae are very abundant. The bacteria are autotrophs that fix CO via the Wood-Ljungdahl pathway and reductive TCA cycles. The genomes encode versatile energy-generating pathways that involve sulfate reduction, hydrogen oxidation and nitrite reduction. Phylogenetic analyses indicate that the organisms are most similar to the isolated magnetotactic bacterium, Candidatus Magnetobacterium bavaricum, with only 89-91% 16S rRNA gene sequence identity. These Nitrospirae bacteria appear to play critical ecosystem roles as primary producers and they are likely central to sulfur cycling in the deep subsurface.
via the Wood-Ljungdahl pathway and reductive TCA cycles. The genomes encode versatile energy-generating pathways that involve sulfate reduction, hydrogen oxidation and nitrite reduction. Phylogenetic analyses indicate that the organisms are most similar to the isolated magnetotactic bacterium, Candidatus Magnetobacterium bavaricum, with only 89-91% 16S rRNA gene sequence identity. These Nitrospirae bacteria appear to play critical ecosystem roles as primary producers and they are likely central to sulfur cycling in the deep subsurface.
天野 由記; Diamond, S.*; Lavy, A.*; Anantharaman, K.*; 宮川 和也; 岩月 輝希; 別部 光里*; 鈴木 庸平*; Thomas, B. C.*; Banfield, J. F.*
no journal, ,
We investigated the microbiology two Japanese subsurface research sites and compared the major groups of organisms lacking cultivated representatives found from other subsurface sites, including a Colorado aquifer and deep aquifers underlying the Crystal Geyser. We analyzed metagenomic data 19 samples from the Horonobe site and 7 from the Mizunami site. DNA sequences from each sample were assembled independently and scaffolds encoding the ribosomal protein S3 sequence were identified. The major characteristic of the microbiology of the Mizumani site that distinguished it from the Horonobe site is local very high abundances of Nitrospirae, Parcubacteria, Ignavibacteria, ANME-2D and Micrarchaeota. In contrast, the Horonobe site has locations that are highly enriched in Altarchiales, Syntrophobacteriales, Atribacteria, ANME-2D and Methanogens. Beyond reshaping the Tree of Life, the societal importance of these discoveries remains little known. However, given the huge inventory of new groups of proteins and pathways in the genomes of these organisms, it is reasonable to anticipate major discoveries will hold relevance, for example, in terms of pharmaceutical discovery. Given the importance of the subsurface as a potential host environment for storage of nuclear waste, finding some commonality would indicate the general relevance of information from one site for prediction of the characteristics of other sites.
天野 由記; Sachdeva, R.*; Gittins, D.*; Anantharaman, K.*; Lei, S.*; 別部 光里*; 望月 陽人; Thomas, B. C.*; 鈴木 庸平*; Banfield, J. F.*
no journal, ,
Underground research laboratories (URLs) provide a window on the deep biosphere and enable investigation of potential microbial impacts on geological disposal, CO and H
and H stored in the subsurface. We carried out the first multi-year study of groundwater microbiomes sampled from defined intervals between 140 and 400 m below the surface of the Horonobe and Mizunami URLs, Japan. We reconstructed draft genomes for
stored in the subsurface. We carried out the first multi-year study of groundwater microbiomes sampled from defined intervals between 140 and 400 m below the surface of the Horonobe and Mizunami URLs, Japan. We reconstructed draft genomes for 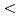 90% of all organisms detected over a four-year period. The Horonobe and Mizunami microbiomes are dissimilar, likely because the Mizunami URL is hosted in granitic rock and the Horonobe URL in sedimentary rock. Despite this, hydrogen metabolism, rubisco-based CO
90% of all organisms detected over a four-year period. The Horonobe and Mizunami microbiomes are dissimilar, likely because the Mizunami URL is hosted in granitic rock and the Horonobe URL in sedimentary rock. Despite this, hydrogen metabolism, rubisco-based CO fixation, reduction of nitrogen compounds and sulfate reduction are well represented functions in microbiomes from both URLs, although methane metabolism is more prevalent at the organic- and CO
fixation, reduction of nitrogen compounds and sulfate reduction are well represented functions in microbiomes from both URLs, although methane metabolism is more prevalent at the organic- and CO -rich Horonobe URL. High fluid flow zones and proximity to subsurface tunnels apparently impact microbial community composition. We detected near-identical genotypes for approximately one third of genomically defined organisms at multiple depths within the Horonobe URL. Sequencing data indicate that at least some of the bacteria are growing, albeit slowly. Genotypically-defined species closely related to those detected in the URLs were identified in three other subsurface environments in the USA. Thus, dispersal rates may be fast enough relative to mutation rates to have limited substantial species divergence. Hydraulic and isotopic measurements predict inter-site movement of groundwater over thousands of years, but it is uncertain whether the permeability is sufficient for transport of microbial cells. Alternatively, Horonobe URL microbiome homogeneity may be explained by dispersal during last glacial period.
-rich Horonobe URL. High fluid flow zones and proximity to subsurface tunnels apparently impact microbial community composition. We detected near-identical genotypes for approximately one third of genomically defined organisms at multiple depths within the Horonobe URL. Sequencing data indicate that at least some of the bacteria are growing, albeit slowly. Genotypically-defined species closely related to those detected in the URLs were identified in three other subsurface environments in the USA. Thus, dispersal rates may be fast enough relative to mutation rates to have limited substantial species divergence. Hydraulic and isotopic measurements predict inter-site movement of groundwater over thousands of years, but it is uncertain whether the permeability is sufficient for transport of microbial cells. Alternatively, Horonobe URL microbiome homogeneity may be explained by dispersal during last glacial period.