Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Arai, Yosuke*; Kuroda, Kenta*; Nomoto, Takuya*; Tin, Z. H.*; Sakuragi, Shunsuke*; Bareille, C.*; Akebi, Shuntaro*; Kurokawa, Kifu*; Kinoshita, Yuto*; Zhang, W.-L.*; et al.
Nature Materials, 21(4), p.410 - 415, 2022/04
Times Cited Count:14 Percentile:75.71(Chemistry, Physical) binding site of a
binding site of a  -lactamase from the halophile by anomalous X-ray diffraction
-lactamase from the halophile by anomalous X-ray diffractionArai, Shigeki; Shibazaki, Chie; Shimizu, Rumi; Adachi, Motoyasu; Tamada, Taro; Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Tokunaga, Masao*; Kuroki, Ryota
Kyushu Shinkurotoronko Kenkyu Senta Nempo, 2014, p.17 - 19, 2016/03
no abstracts in English
Mori, Masanobu*; Sagara, Katsuya*; Arai, Kaori*; Nakatani, Nobutake*; Ohira, Shinichi*; Toda, Kei*; Itabashi, Hideyuki*; Kozaki, Daisuke*; Sugo, Yumi; Watanabe, Shigeki; et al.
Journal of Chromatography A, 1431, p.131 - 137, 2016/01
Times Cited Count:10 Percentile:38.40(Biochemical Research Methods) sp. AS-131 isolated from Antarctic Ocean
sp. AS-131 isolated from Antarctic OceanYonezawa, Yasushi*; Nagayama, Aiko*; Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Arai, Shigeki; Kuroki, Ryota; Watanabe, Keiichi*; Arakawa, Tsutomu*; Tokunaga, Masao*
Protein Journal, 34(4), p.275 - 283, 2015/08
Times Cited Count:4 Percentile:10.01(Biochemistry & Molecular Biology)Nucleoside diphosphate kinase isolated from psychrophilic  sp. AS-131 (ASNDK) was expressed in
sp. AS-131 (ASNDK) was expressed in  and purified to homogeneity. Comparing to mesophilic NDK isolated from
and purified to homogeneity. Comparing to mesophilic NDK isolated from  , ASNDK exhibited highly elevated thermolability: (1)
, ASNDK exhibited highly elevated thermolability: (1) 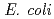 expression at 37
expression at 37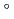 C as a denatured insoluble form, and (2) 30
C as a denatured insoluble form, and (2) 30 C lower optimum temperature of enzymatic activity. The subunit structure of ASNDK was suggested to be dimer, as in NDKs isolated from moderate halophiles.
C lower optimum temperature of enzymatic activity. The subunit structure of ASNDK was suggested to be dimer, as in NDKs isolated from moderate halophiles.
 -lactamase from the moderate halophile
-lactamase from the moderate halophile 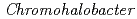 sp.560 and the discovery of a Cs
sp.560 and the discovery of a Cs -selective binding site
-selective binding siteArai, Shigeki; Yonezawa, Yasushi*; Okazaki, Nobuo*; Matsumoto, Fumiko*; Shibazaki, Chie; Shimizu, Rumi; Yamada, Mitsugu*; Adachi, Motoyasu; Tamada, Taro; Kawamoto, Masahide*; et al.
Acta Crystallographica Section D, 71(3), p.541 - 554, 2015/03
Times Cited Count:8 Percentile:52.61(Biochemical Research Methods)The crystal structure of halophilic  -lactamase from
-lactamase from 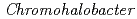 sp.560 (HaBLA) was determined using X-ray crystallography. Moreover, the locations of bound Sr
sp.560 (HaBLA) was determined using X-ray crystallography. Moreover, the locations of bound Sr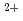 and Cs
and Cs ions were identified by anomalous X-ray diffraction. The location of one Cs
ions were identified by anomalous X-ray diffraction. The location of one Cs specific binding site was identified on HaBLA even in the presence of 9-fold molar excess of Na
specific binding site was identified on HaBLA even in the presence of 9-fold molar excess of Na (90 mM Na
(90 mM Na /10 mM Cs
/10 mM Cs ). This Cs
). This Cs binding site is formed by two main-chain O atoms and an aromatic ring of a side chain of Trp. An aromatic ring of Trp interacts with Cs
binding site is formed by two main-chain O atoms and an aromatic ring of a side chain of Trp. An aromatic ring of Trp interacts with Cs by the cation-
by the cation-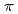 interaction. The observation of a selective and high-affinity Cs
interaction. The observation of a selective and high-affinity Cs binding site provides important information that is useful for designing artificial Cs
binding site provides important information that is useful for designing artificial Cs binding sites useful in bioremediation of radioactive isotopes.
binding sites useful in bioremediation of radioactive isotopes.
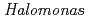 sp.593
sp.593Arai, Shigeki; Yonezawa, Yasushi*; Ishibashi, Matsujiro*; Matsumoto, Fumiko*; Adachi, Motoyasu; Tamada, Taro; Tokunaga, Hiroko*; Blaber, M.; Tokunaga, Masao*; Kuroki, Ryota
Acta Crystallographica Section D, 70(3), p.811 - 820, 2014/03
Times Cited Count:13 Percentile:63.53(Biochemical Research Methods)In order to clarify the structural basis of halophilic characteristics of an alkaline phosphatase derived from the moderate halophile 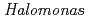 sp.593 (HaAP), the tertiary structure of HaAP was determined to 2.1
sp.593 (HaAP), the tertiary structure of HaAP was determined to 2.1 resolution by X-ray crystallography. Structural properties of surface negative charge and core hydrophobicity are shown to be intermediate between halophile and non-halophile characteristics, and may explain the unique functional adaptation to a wide-range of salt concentration.
resolution by X-ray crystallography. Structural properties of surface negative charge and core hydrophobicity are shown to be intermediate between halophile and non-halophile characteristics, and may explain the unique functional adaptation to a wide-range of salt concentration.
Adachi, Motoyasu; Arai, Shigeki; Hiromoto, Takeshi; Kuroki, Ryota
Hamon, 24(1), p.45 - 49, 2014/02
Protein structure analysis using neutron diffraction (neutron protein crystallography; NPC) is gaining greater importance in the understanding of structure and function relationships of biological macromolecules such as proteins and DNA. Current developments of neutron diffractometers installed at the JAEA research reactor and pulsed neutron source permit observation of the locations of hydrogen atoms and hydrating water molecules and help understanding of important mechanisms of chemical reactions catalyzed by biological macromolecules. Here, we introduce practical approaches of NPC including sample preparation, crystal growth, structure determination and utilization of information obtained from NPC.
Saegusa, Jun; Kurikami, Hiroshi; Yasuda, Ryo; Kurihara, Kazuo; Arai, Shigeki; Kuroki, Ryota; Matsuhashi, Shimpei; Ozawa, Takashi; Goto, Hiroaki; Takano, Takao; et al.
Health Physics, 104(3), p.243 - 250, 2013/03
Times Cited Count:3 Percentile:24.26(Environmental Sciences)After the Nuclear accident on March 2011, water discharge from many outdoor swimming pools in the Fukushima prefecture was suspended out of concern that radiocesium in the pool water would flow into farmlands. We have reviewed the existing flocculation method for decontaminating pool water and established a practical decontamination method by demonstrating the process at several pools in the Fukushima prefecture.
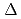 1-tetrahydrocannabinolic acid (THCA) synthase, the enzyme controlling the psychoactivity of
1-tetrahydrocannabinolic acid (THCA) synthase, the enzyme controlling the psychoactivity of 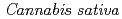
Shoyama, Yoshinari*; Tamada, Taro; Kurihara, Kazuo; Takeuchi, Ayako*; Taura, Futoshi*; Arai, Shigeki; Blaber, M.*; Shoyama, Yukihiro*; Morimoto, Satoshi*; Kuroki, Ryota
Journal of Molecular Biology, 423(1), p.96 - 105, 2012/10
Times Cited Count:90 Percentile:90.02(Biochemistry & Molecular Biology)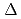 1-tetrahydrocannabinolic acid (THCA) synthase catalyzes the oxidative cyclization of cannabigerolic acid (CBGA) into THCA, the precursor of the primary psychoactive agent
1-tetrahydrocannabinolic acid (THCA) synthase catalyzes the oxidative cyclization of cannabigerolic acid (CBGA) into THCA, the precursor of the primary psychoactive agent  1-tetrahydrocannabinol in
1-tetrahydrocannabinol in 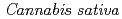 . The structure-function relationship of THCA synthase was investigated by X-ray structure determination (2.75
. The structure-function relationship of THCA synthase was investigated by X-ray structure determination (2.75  resolution) and mutational analysis. Specific amino acid residues were identified in the active site of THCA synthase that are involved in the oxidative cyclization of the CBGA substrate.
resolution) and mutational analysis. Specific amino acid residues were identified in the active site of THCA synthase that are involved in the oxidative cyclization of the CBGA substrate.
Ishibashi, Matsujiro*; Uchino, Manami*; Arai, Shigeki; Kuroki, Ryota; Arakawa, Tsutomu*; Tokunaga, Masao*
Archives of Biochemistry and Biophysics, 525(1), p.47 - 52, 2012/09
Times Cited Count:7 Percentile:19.87(Biochemistry & Molecular Biology)Nucleoside diphosphate kinase (HsNDK) from Halobacterium salinarum requires salt at high concentrations for folding. A D148C mutant, in which Asp148 was replaced with Cys, was designed to enhance stability and folding in low salt solution by S-S bond. It showed increased thermal stability by about 10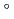 C in 0.2 M NaCl over the wild type HsNDK. It refolded from heat-denaturation even in 0.1 M NaCl, while the wild type required 2 M NaCl to achieve the same level of activity recovery. This enhanced refolding is due to the three S-S bonds between two basic dimeric units in the hexameric HsNDK structure. Moreover, salt concentration dependency of heat-denaturation process and refolding process of the wild type and D148C mutant HsNDKs were investigated by the circular dichroism and native-PAGE analysis.
C in 0.2 M NaCl over the wild type HsNDK. It refolded from heat-denaturation even in 0.1 M NaCl, while the wild type required 2 M NaCl to achieve the same level of activity recovery. This enhanced refolding is due to the three S-S bonds between two basic dimeric units in the hexameric HsNDK structure. Moreover, salt concentration dependency of heat-denaturation process and refolding process of the wild type and D148C mutant HsNDKs were investigated by the circular dichroism and native-PAGE analysis.
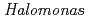 sp. nucleoside diphosphate kinase
sp. nucleoside diphosphate kinaseArai, Shigeki; Yonezawa, Yasushi; Okazaki, Nobuo; Matsumoto, Fumiko; Tamada, Taro; Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Blaber, M.; Tokunaga, Masao*; Kuroki, Ryota
Protein Science, 21(4), p.498 - 510, 2012/04
Times Cited Count:15 Percentile:34.92(Biochemistry & Molecular Biology)In order to clarify the oligomer state of nucleoside diphosphate kinase (NDK) from moderately halophilic  sp. 593 (HaNDK), the crystal structure of HaNDK was determined by X-ray crystallography. The crystal structures of the wild-type HaNDK and the mutant HaNDK (E134A) showed a dimer and a tetramer, respectively. The higher ordered association of proteins usually contributes to an increase in thermal stability and substrate affinity. The change in the assembly form by a minimum mutation may be an effective way for NDK to acquire molecular characteristics suited to various circumstances.
sp. 593 (HaNDK), the crystal structure of HaNDK was determined by X-ray crystallography. The crystal structures of the wild-type HaNDK and the mutant HaNDK (E134A) showed a dimer and a tetramer, respectively. The higher ordered association of proteins usually contributes to an increase in thermal stability and substrate affinity. The change in the assembly form by a minimum mutation may be an effective way for NDK to acquire molecular characteristics suited to various circumstances.
 DSM3043; Both residues 134 and 136 are critical for the tetramer assembly
DSM3043; Both residues 134 and 136 are critical for the tetramer assemblyTokunaga, Hiroko*; Izutsu, Kenichi*; Arai, Shigeki; Yonezawa, Yasushi; Kuroki, Ryota; Arakawa, Tsutomu*; Tokunaga, Masao*
Enzyme and Microbial Technology, 46(2), p.129 - 135, 2010/02
Times Cited Count:6 Percentile:20.05(Biotechnology & Applied Microbiology)Both wild-type nucleoside diphosphate kinase from moderately halophilc  (CsNDK (GNE), GNE represents Gly134-Asn135-Glu136) and mutant CsNDK (ANE), both of which have a neutral amino acid at residue 134, were found to form a dimer. These constructs contain Glu136, which may also cause steric barrier and charge repulsion. A double mutant, CsNDK (ANT), having Thr at 136 resulted in stable tetrameric assembly, supporting the above notion. A mutant CsNDK (GNT) reverted, however, to a dimer again, indicating that the introduced Ala residue at 134th in the double mutant generated a hydrophobic cluster consisting of the Ala residues and thereby stabilized dimer-dimer association of CsNDK assembly, while Gly destabilized it due to the loss of this cluster. Based on these observations, it is evident that both residues 134 and 136 contribute to the subunit assembly of CsNDK.
(CsNDK (GNE), GNE represents Gly134-Asn135-Glu136) and mutant CsNDK (ANE), both of which have a neutral amino acid at residue 134, were found to form a dimer. These constructs contain Glu136, which may also cause steric barrier and charge repulsion. A double mutant, CsNDK (ANT), having Thr at 136 resulted in stable tetrameric assembly, supporting the above notion. A mutant CsNDK (GNT) reverted, however, to a dimer again, indicating that the introduced Ala residue at 134th in the double mutant generated a hydrophobic cluster consisting of the Ala residues and thereby stabilized dimer-dimer association of CsNDK assembly, while Gly destabilized it due to the loss of this cluster. Based on these observations, it is evident that both residues 134 and 136 contribute to the subunit assembly of CsNDK.
Adachi, Motoyasu; Ohara, Takashi; Kurihara, Kazuo; Tamada, Taro; Honjo, Eijiro; Okazaki, Nobuo; Arai, Shigeki; Shoyama, Yoshinari; Kimura, Kaname*; Matsumura, Hiroyoshi*; et al.
Proceedings of the National Academy of Sciences of the United States of America, 106(12), p.4641 - 4646, 2009/03
Times Cited Count:115 Percentile:90.49(Multidisciplinary Sciences)To further understand the catalytic mechanism and inhibitor recognition of HIV-1 protease, we need to determine the locations of key hydrogen atoms in the catalytic aspartates Asp25 and Asp125. The structure of HIV-1 protease in complex with transition-state analog KNI-272 was determined by combined neutron crystallography at 1.9  resolution and X-ray crystallography at 1.4
resolution and X-ray crystallography at 1.4  resolution. The resulting structural data shows that the catalytic residue Asp25 is protonated and that Asp125 is deprotonated. The proton on Asp25 makes a hydrogen bond with the carbonyl group of the allophenylnorstatine group in KNI-272. The deprotonated Asp125 bonds to the hydroxyl proton of Apns. The results provide direct experimental evidence for proposed aspects of the catalytic mechanism of HIV-1 protease; and can therefore contribute substantially to the development of specific inhibitors for therapeutic application.
resolution. The resulting structural data shows that the catalytic residue Asp25 is protonated and that Asp125 is deprotonated. The proton on Asp25 makes a hydrogen bond with the carbonyl group of the allophenylnorstatine group in KNI-272. The deprotonated Asp125 bonds to the hydroxyl proton of Apns. The results provide direct experimental evidence for proposed aspects of the catalytic mechanism of HIV-1 protease; and can therefore contribute substantially to the development of specific inhibitors for therapeutic application.
Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Arisaka, Fimio*; Arai, Shigeki; Kuroki, Ryota; Arakawa, Tsutomu*; Tokunaga, Masao*
FEBS Letters, 582(7), p.1049 - 1054, 2008/04
Times Cited Count:17 Percentile:39.41(Biochemistry & Molecular Biology) nucleoside diphosphate kinase (HaNDK) forms a dimeric assembly and
nucleoside diphosphate kinase (HaNDK) forms a dimeric assembly and 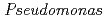 NDK (PaNDK) forms a tetrameric assembly. The mutation of Glu134 to Ala in HaNDK resulted in conversion of the native dimeric structure to the tetramer assembly. Conversely, the mutation of Ala134 to Glu in PaNDK leads to conversion from tetramer to dimer assembly, indicating that a single amino acid substitution at position 134 results in an alteration of the oligomeric structure of NDK. Modeling structure of HaNDK and PaNDK, based on the crystal structure of
NDK (PaNDK) forms a tetrameric assembly. The mutation of Glu134 to Ala in HaNDK resulted in conversion of the native dimeric structure to the tetramer assembly. Conversely, the mutation of Ala134 to Glu in PaNDK leads to conversion from tetramer to dimer assembly, indicating that a single amino acid substitution at position 134 results in an alteration of the oligomeric structure of NDK. Modeling structure of HaNDK and PaNDK, based on the crystal structure of 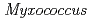 NDK, suggested sufficient repulsion by Glu134 to disrupt dimer-dimer interaction to form tetramer.
NDK, suggested sufficient repulsion by Glu134 to disrupt dimer-dimer interaction to form tetramer.
Arai, Shigeki; Chatake, Toshiyuki*; Niimura, Nobuo
Nihon Kessho Gakkai-Shi, 48(2), p.133 - 139, 2006/04
It has long been suspected that the structure and function of a DNA duplex can be strongly dependent on its degree of hydration. By neutron diffraction experiments, we have succeeded in determining most of the hydrogen (H) and deuterium (D) atomic positions in the d(CCATTAATGG) duplex. Moreover, the D positions in 27 D
duplex. Moreover, the D positions in 27 D O molecules have been determined. In particular, the complex water network in the minor groove has been observed in detail. By a combined structural analysis using 2.0 AA resolution X-ray and 3.0 AA resolution neutron data, it is clear that the spine of hydration is built up, not only by a simple hexagonal hydration pattern (as reported in prior X-ray studies), but also by many other water bridges hydrogen-bonded to the DNA strands. The complexity of the hydration pattern in the minor groove is derived from an extraordinary variety of orientations displayed by the water molecules.
O molecules have been determined. In particular, the complex water network in the minor groove has been observed in detail. By a combined structural analysis using 2.0 AA resolution X-ray and 3.0 AA resolution neutron data, it is clear that the spine of hydration is built up, not only by a simple hexagonal hydration pattern (as reported in prior X-ray studies), but also by many other water bridges hydrogen-bonded to the DNA strands. The complexity of the hydration pattern in the minor groove is derived from an extraordinary variety of orientations displayed by the water molecules.
Niimura, Nobuo; Arai, Shigeki; Kurihara, Kazuo; Chatake, Toshiyuki*; Tanaka, Ichiro*; Bau, R.*
Cellular and Molecular Life Sciences, 63(3), p.285 - 300, 2006/02
Times Cited Count:41 Percentile:36.92(Biochemistry & Molecular Biology)Neutron diffraction provides an experimental method of directly locating hydrogen atoms in proteins and DNA oligomers. Three different types of high resolution neutron diffractometers for biological macromolecules have been constructed in Japan, France and the U.S.A., and they have all been actively used in recent years to determine the crystal structures of numerous proteins. Examples include the detailed geometries of hydrogen bonds, information on H/D exchange in proteins, the unambiguous location of protons, the role of key hydrogen atoms in enzymatic activity and thermostability, and the dynamical behavior of hydration structures, all of which have been extracted from these structural results and reviewed in this article. Other important techniques, such as the optimization of growth of large single crystals using phase diagrams, the preparation of fully deuterated proteins, the introduction of cryogenic techniques to neutron protein crystallography, and the establishment of a "hydrogen and hydration in proteins" database, will also be described in this paper.
 observed by neutron diffraction measurements
observed by neutron diffraction measurementsArai, Shigeki; Chatake, Toshiyuki*; Ohara, Takashi; Kurihara, Kazuo; Tanaka, Ichiro*; Suzuki, Nobuhiro*; Fujimoto, Zui*; Mizuno, Hiroshi*; Niimura, Nobuo
Nucleic Acids Research, 33(9), p.3017 - 3024, 2005/05
Times Cited Count:94 Percentile:82.51(Biochemistry & Molecular Biology)It has long been suspected that the structure and function of a DNA duplex can be strongly dependent on its degree of hydration. By neutron diffraction experiments, we have succeeded in determining most of the hydrogen (H) and deuterium (D) atomic positions in the d(CCATTAATGG) duplex. Moreover, the D positions in 27 D
duplex. Moreover, the D positions in 27 D O molecules have been determined. In particular, the complex water network in the minor groove has been observed in detail. By a combined structural analysis using 2.0 Å resolution X-ray and 3.0 Å resolution neutron data, it is clear that the spine of hydration is built up, not only by a simple hexagonal hydration pattern (as reported in prior X-ray studies), but also by many other water bridges hydrogen-bonded to the DNA strands. The complexity of the hydration pattern in the minor groove is derived from an extraordinary variety of orientations displayed by the water molecules.
O molecules have been determined. In particular, the complex water network in the minor groove has been observed in detail. By a combined structural analysis using 2.0 Å resolution X-ray and 3.0 Å resolution neutron data, it is clear that the spine of hydration is built up, not only by a simple hexagonal hydration pattern (as reported in prior X-ray studies), but also by many other water bridges hydrogen-bonded to the DNA strands. The complexity of the hydration pattern in the minor groove is derived from an extraordinary variety of orientations displayed by the water molecules.
Niimura, Nobuo; Arai, Shigeki; Kurihara, Kazuo; Chatake, Toshiyuki*; Tanaka, Ichiro*; Bau, R.*
Hydrogen- and Hydration-Sensitive Structural Biology, p.17 - 35, 2005/00
At the JAERI, we have constructed several high-resolution neutron diffractometers dedicated to biological macromolecules (called BIX-type diffractometers), which use a monochromatized neutron beam and a neutron imaging plate detector. In this paper, we review several interesting results regarding hydrogen positions and hydration in proteins, obtained using the two BIX-type diffractometers in JAERI. The general subject of neutron protein crystallography has been reviewed by several authors, and several selected topics have been discussed.
Arai, Shigeki; Chatake, Toshiyuki; Suzuki, Nobuhiro*; Mizuno, Hiroshi*; Niimura, Nobuo
Acta Crystallographica Section D, 60(6), p.1032 - 1039, 2004/06
Times Cited Count:17 Percentile:75.55(Biochemical Research Methods)The parameters used for evaluating biomacromolecular crystal quality (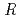
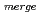 ,
, 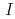 /
/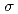 (
(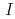 ), maximum resolution and mosaicity) strongly depend on the diffraction experimental conditions. In this paper we describe the distinctive features of the relative Wilson plot method, and we show that the overall B-factor obtained from this plot is given as a more appropriate to characterize protein crystals. The relative Wilson plot has been applied to the characterization of crystals of a B-DNA decamer d(CCATTAATGG), and crystals of the proteins DsrD (dissimilatory sulfite reductase D) and hen egg-white lysozyme (HEWL) which we have studied by neutron diffraction. We have found that the crystal qualities of the B-DNA decamer and DsrD significantly depend on the regions of the crystallization phase diagram from which samples were taken. However, in the case of HEWL, crystal quality appears to be independent on the region of the crystallization phase diagram.
), maximum resolution and mosaicity) strongly depend on the diffraction experimental conditions. In this paper we describe the distinctive features of the relative Wilson plot method, and we show that the overall B-factor obtained from this plot is given as a more appropriate to characterize protein crystals. The relative Wilson plot has been applied to the characterization of crystals of a B-DNA decamer d(CCATTAATGG), and crystals of the proteins DsrD (dissimilatory sulfite reductase D) and hen egg-white lysozyme (HEWL) which we have studied by neutron diffraction. We have found that the crystal qualities of the B-DNA decamer and DsrD significantly depend on the regions of the crystallization phase diagram from which samples were taken. However, in the case of HEWL, crystal quality appears to be independent on the region of the crystallization phase diagram.
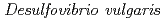
Chatake, Toshiyuki; Mizuno, Nobuhiro*; Voordown, G.*; Higuchi, Yoshiki*; Arai, Shigeki; Tanaka, Ichiro; Niimura, Nobuo
Acta Crystallographica Section D, 59(Part2), p.2306 - 2309, 2003/12
Times Cited Count:19 Percentile:75.12(Biochemical Research Methods)Issimilatory sulfite reductase D (DsrD) from 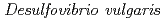 has been crystallized for a neutron diffraction study. The initial crystals obtained were too small for the neutron experiment. In order to obtain a larger crystal (1 mm
has been crystallized for a neutron diffraction study. The initial crystals obtained were too small for the neutron experiment. In order to obtain a larger crystal (1 mm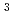 ), a combination of two techniques was used to find the optimum crystallization conditions: a crystallization phase diagram, followed by crystal-quality assessment via X-ray diffraction. Using conditions determined in this manner, a large single crystal (1.7 mm
), a combination of two techniques was used to find the optimum crystallization conditions: a crystallization phase diagram, followed by crystal-quality assessment via X-ray diffraction. Using conditions determined in this manner, a large single crystal (1.7 mm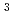 ) of the DsrD protein was subsequently grown in D
) of the DsrD protein was subsequently grown in D O solution by the macro-seeding technique. The neutron diffraction experiment was carried out using the BIX-3 diffractometer at the Japan Atomic Energy Research Institute (JAERI), and the diffraction data up to 2.4 AA resolution could be collected from this crystal.
O solution by the macro-seeding technique. The neutron diffraction experiment was carried out using the BIX-3 diffractometer at the Japan Atomic Energy Research Institute (JAERI), and the diffraction data up to 2.4 AA resolution could be collected from this crystal.