Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Hiromoto, Takeshi; Adachi, Motoyasu; Shibazaki, Chie; Schrader, T. E.*; Ostermann, A.*; Kuroki, Ryota
JPS Conference Proceedings (Internet), 8, p.033003_1 - 033003_6, 2015/09
T4 phage lysozyme (T4L) is an endoacetylmuramidase that degrades the murein of the bacterial cell wall by cleaving the  -1,4-glycosidic bond between
-1,4-glycosidic bond between 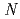 -acetylmuramic acid and
-acetylmuramic acid and 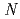 -acetylglucosamine. We previously reported that the substitution of the catalytic Thr26 with the nucleophilic His converts the wild-type (WT) T4L from an inverting to a retaining glycosidase, in which the
-acetylglucosamine. We previously reported that the substitution of the catalytic Thr26 with the nucleophilic His converts the wild-type (WT) T4L from an inverting to a retaining glycosidase, in which the  -configuration of the substrate is retained in the product. We also found that the Thr26His (T26H) mutant can catalyze a transglycosylation reaction more effectively than hydrolysis, although the WT-T4L has no transglycosidase activity. To clarify the role of the substituted His26 in transglycosylation and to investigate its relationship to the neighboring acidic residue Asp20 using neutron crystallography, a perdeuterated recombinant protein of the T26H mutant (d-T26H) was prepared for crystallization. The perdeuterated form was produced in
-configuration of the substrate is retained in the product. We also found that the Thr26His (T26H) mutant can catalyze a transglycosylation reaction more effectively than hydrolysis, although the WT-T4L has no transglycosidase activity. To clarify the role of the substituted His26 in transglycosylation and to investigate its relationship to the neighboring acidic residue Asp20 using neutron crystallography, a perdeuterated recombinant protein of the T26H mutant (d-T26H) was prepared for crystallization. The perdeuterated form was produced in 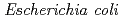 cells cultured in deuterated-rich media. After purification, the d-T26H mutant was crystallized under deuterated conditions and grown to a volume of 0.12 mm
cells cultured in deuterated-rich media. After purification, the d-T26H mutant was crystallized under deuterated conditions and grown to a volume of 0.12 mm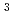 using a macroseeding technique. A preliminary neutron-diffraction experiment at 100 K at the FRM II research reactor (Munich, Germany) gave diffraction spots of up to 2.8
using a macroseeding technique. A preliminary neutron-diffraction experiment at 100 K at the FRM II research reactor (Munich, Germany) gave diffraction spots of up to 2.8  resolution after a 1.5 h exposure.
resolution after a 1.5 h exposure.
Niimura, Nobuo; Chatake, Toshiyuki; Ostermann, A.; Kurihara, Kazuo; Tanaka, Ichiro
Zeitschrift f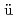 r Kristallographie, 218(2), p.96 - 107, 2003/03
r Kristallographie, 218(2), p.96 - 107, 2003/03
no abstracts in English
Chatake, Toshiyuki; Ostermann, A.; Kurihara, Kazuo; Parak, F.*; Niimura, Nobuo
Proteins: Structure, Function, and Bioinformatics, 50(3), p.516 - 523, 2003/02
Times Cited Count:54 Percentile:76.77(Biochemistry & Molecular Biology)no abstracts in English
Niimura, Nobuo*; Tanaka, Ichiro*; Onishi, Yuki*; Ostermann, A.*; Kurihara, Kazuo; Honjo, Eijiro; Kuroki, Ryota; Futami, Junichiro*; Yamada, Hidenori*
no journal, ,
Our goal is to develop new methods for determining macromolecular crystal structures 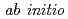 from neutron diffraction data and to validate these methods by making applications to several proteins. The proposed methods, which have the potential to be more powerful than X-ray ones, exploit perfectly isomorphous pairs of crystals that differ by replacement of D atoms with H atoms in selected amino-acid residues. This research involves collaboration among teams on three continents that have expertise in different aspects of neutron crystallography. The Japanese team measures high-resolution neutron diffraction data from protein crystals to answer questions about H bonding, protonation, hydration, and enzyme mechanisms. The team will focus on fundamental and simple proteins such as myoglobin, insulin, and RNase A at the first stage and then on rubredoxin, aldose reductase and other samples from the French team. In the meeting the research plan of the Japanese team will be reported.
from neutron diffraction data and to validate these methods by making applications to several proteins. The proposed methods, which have the potential to be more powerful than X-ray ones, exploit perfectly isomorphous pairs of crystals that differ by replacement of D atoms with H atoms in selected amino-acid residues. This research involves collaboration among teams on three continents that have expertise in different aspects of neutron crystallography. The Japanese team measures high-resolution neutron diffraction data from protein crystals to answer questions about H bonding, protonation, hydration, and enzyme mechanisms. The team will focus on fundamental and simple proteins such as myoglobin, insulin, and RNase A at the first stage and then on rubredoxin, aldose reductase and other samples from the French team. In the meeting the research plan of the Japanese team will be reported.