Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
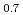 La
La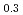 B
B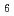 studied by resonant X-ray diffraction
studied by resonant X-ray diffractionMatsumura, Takeshi*; Michimura, Shinji*; Inami, Toshiya; Otsubo, Toru*; Tanida, Hiroshi*; Iga, Fumitoshi*; Sera, Masafumi*
JPS Conference Proceedings (Internet), 3, p.014008_1 - 014008_6, 2014/06
We study the 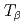 -type antiferro-octupole (AFO) ordered state of Ce
-type antiferro-octupole (AFO) ordered state of Ce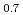 La
La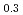 B
B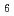 below 1.5 K by resonant X-ray diffraction in magnetic fields. Multipole moments induced by the field in the AFO phase are identified. In a mean-field model for the
below 1.5 K by resonant X-ray diffraction in magnetic fields. Multipole moments induced by the field in the AFO phase are identified. In a mean-field model for the 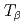 AFO order within the
AFO order within the 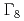 quartet crystal-field ground state, the
quartet crystal-field ground state, the 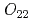 and
and 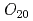 antiferroquadrupoles (AFQ) are expected as mostly induced AFQ. However, in contrast to this expectation, the main induced moment is found to be the
antiferroquadrupoles (AFQ) are expected as mostly induced AFQ. However, in contrast to this expectation, the main induced moment is found to be the 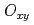 -type AFQ.
-type AFQ.
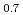 La
La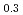 B
B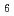 from resonant X-ray diffraction in magnetic fields
from resonant X-ray diffraction in magnetic fieldsMatsumura, Takeshi*; Michimura, Shinji*; Inami, Toshiya; Otsubo, Toru*; Tanida, Hiroshi*; Iga, Fumitoshi*; Sera, Masafumi*
Physical Review B, 89(1), p.014422_1 - 014422_13, 2014/01
Times Cited Count:14 Percentile:49.64(Materials Science, Multidisciplinary)The multipole ordered phase in Ce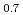 La
La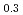 B
B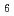 (phase IV) has been studied by resonant X-ray diffraction in magnetic fields. By utilizing diamond X-ray phase plates to rotate the incident linear polarization and a conventional crystal analyzer system, full linear polarization analysis has been performed to identify the order parameters. The analysis shows that the
(phase IV) has been studied by resonant X-ray diffraction in magnetic fields. By utilizing diamond X-ray phase plates to rotate the incident linear polarization and a conventional crystal analyzer system, full linear polarization analysis has been performed to identify the order parameters. The analysis shows that the 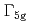 quadrupoles are more induced by the field than the
quadrupoles are more induced by the field than the 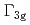 quadrupoles in phase IV, in disagreement with mean-field calculations, in which the
quadrupoles in phase IV, in disagreement with mean-field calculations, in which the 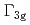 quadrupole is the main induced moment. Hence, we consider that large fluctuations of the
quadrupole is the main induced moment. Hence, we consider that large fluctuations of the 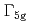 quadrupole is hidden behind the primary ordering of the octupole.
quadrupole is hidden behind the primary ordering of the octupole.
Yamasaki, Chisato*; Murakami, Katsuhiko*; Fujii, Yasuyuki*; Sato, Yoshiharu*; Harada, Erimi*; Takeda, Junichi*; Taniya, Takayuki*; Sakate, Ryuichi*; Kikugawa, Shingo*; Shimada, Makoto*; et al.
Nucleic Acids Research, 36(Database), p.D793 - D799, 2008/01
Times Cited Count:52 Percentile:70.47(Biochemistry & Molecular Biology)Here we report the new features and improvements in our latest release of the H-Invitational Database, a comprehensive annotation resource for human genes and transcripts. H-InvDB, originally developed as an integrated database of the human transcriptome based on extensive annotation of large sets of fulllength cDNA (FLcDNA) clones, now provides annotation for 120 558 human mRNAs extracted from the International Nucleotide Sequence Databases (INSD), in addition to 54 978 human FLcDNAs, in the latest release H-InvDB. We mapped those human transcripts onto the human genome sequences (NCBI build 36.1) and determined 34 699 human gene clusters, which could define 34 057 protein-coding and 642 non-protein-coding loci; 858 transcribed loci overlapped with predicted pseudogenes.
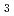 S
S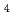
Michimura, Shinji; Inami, Toshiya; Otsubo, Toru*; Matsumura, Takeshi*; Tanida, Hiroshi*; Sera, Masafumi*; Matsuoka, Eiichi*; Watahiki, Masanori*; Tanigaki, Katsumi*; Onodera, Hideya*
no journal, ,
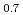 La
La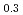 B
B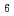 in magnetic field
in magnetic fieldMichimura, Shinji; Inami, Toshiya; Otsubo, Toru*; Matsumura, Takeshi*; Tanida, Hiroshi*; Sera, Masafumi*; Iga, Fumitoshi*
no journal, ,
no abstracts in English
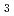 S
S by a resonant X-ray diffraction
by a resonant X-ray diffractionMichimura, Shinji; Inami, Toshiya; Otsubo, Toru*; Matsumura, Takeshi*; Tanida, Hiroshi*; Sera, Masafumi*; Matsuoka, Eiichi*; Watahiki, Masanori*; Tanigaki, Katsumi*; Onodera, Hideya*
no journal, ,
no abstracts in English