Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Yamasaki, Satoshi*; Terada, Toru*; Kono, Hidetoshi; Shimizu, Kentaro*; Sarai, Akinori*
Nucleic Acids Research, 40(17), p.e129_1 - e129_7, 2012/09
Times Cited Count:20 Percentile:44.47(Biochemistry & Molecular Biology)Yamasaki, Satoshi*; Terada, Toru*; Shimizu, Kentaro*; Kono, Hidetoshi; Sarai, Akinori*
Nucleic Acids Research, 37(20), p.e135_1 - e135_9, 2009/09
Times Cited Count:7 Percentile:14.52(Biochemistry & Molecular Biology)Sarai, Akinori*; Kono, Hidetoshi
Jikken Igaku, 26(7), p.1099 - 1105, 2008/04
no abstracts in English
Fujii, Satoshi*; Kono, Hidetoshi; Takenaka, Shigeori*; Go, Nobuhiro; Sarai, Akinori*
Nucleic Acids Research, 35(18), p.6063 - 6074, 2007/09
Times Cited Count:106 Percentile:87.05(Biochemistry & Molecular Biology)Proteins recognize specific DNA sequences not only through direct contact between amino acids and bases, but also indirectly based on the sequence-dependent conformation and deformability of the DNA (indirect readout). We used molecular dynamics simulations to analyze the sequence-dependent DNA conformations of all 136 possible tetrameric sequences sandwiched between CGCG sequences. The deformability of dimeric steps obtained by the simulations is consistent with that by the crystal structures. The simulation results further showed that the conformation and deformability of the tetramers can highly depend on the flanking base-pairs. The conformations of xATx tetramers show the most rigidity and are not affected by the flanking base-pairs and the xYRx show by contrast the greatest flexibility and change their conformations depending on the base-pairs at both ends, suggesting tetramers with the same central dimer can show different deformabilities. These results suggest that analysis of dimeric steps alone may overlook some conformational features of DNA and provide insight into the mechanism of indirect readout during protein-DNA recognition. Moreover, the sequence dependence of DNA conformation and deformability may be used to estimate the contribution of indirect readout to the specificity of protein-DNA recognition as well as nucleosome positioning and large-scale behavior of nucleic acids.
Sarai, Akinori*; Kono, Hidetoshi
Jikken Igaku, 25(10), p.77 - 84, 2007/06
no abstracts in English
Yonetani, Yoshiteru*; Kono, Hidetoshi; Fujii, Satoshi*; Sarai, Akinori*; Go, Nobuhiro
Molecular Simulation, 33(1-2), p.103 - 107, 2007/01
Times Cited Count:5 Percentile:16.09(Chemistry, Physical)DNA tetramer sequences AATT and TTAA are known to be conformationally more rigid and flexible, respectively. In this study, we carry out molecular dynamics simulations of these two sequences, and investigate the characteristic hydration pattern. The rigid AATT is found to be more likely to construct the hydration spine in the minor groove than the flexible TTAA. The result suggests that the hydration water molecules play a critical role for determining the sequence dependent deformability of DNA conformation.
Ahmad, S.*; Kono, Hidetoshi; Ara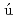 zo-Bravo, M. J.*; Sarai, Akinori*
zo-Bravo, M. J.*; Sarai, Akinori*
Nucleic Acids Research, 34(Suppl.2), p.W124 - W127, 2006/07
Times Cited Count:31 Percentile:48.89(Biochemistry & Molecular Biology)no abstracts in English
Gromiha, M. M.*; Siebers, J. G.*; Selvaraj, S.*; Kono, Hidetoshi; Sarai, Akinori*
Gene, 364, p.108 - 113, 2005/12
Times Cited Count:28 Percentile:45.90(Genetics & Heredity)no abstracts in English
Ara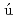 zo-Bravo, M. J.*; Fujii, Satoshi*; Kono, Hidetoshi; Ahmad, S.*; Sarai, Akinori*
zo-Bravo, M. J.*; Fujii, Satoshi*; Kono, Hidetoshi; Ahmad, S.*; Sarai, Akinori*
Journal of the American Chemical Society, 127(46), p.16074 - 16089, 2005/11
Times Cited Count:46 Percentile:72.44(Chemistry, Multidisciplinary)no abstracts in English
Sarai, Akinori*; Kono, Hidetoshi
Annual Review of Biophysics and Biomolecular Structure, 34, p.379 - 398, 2005/06
Times Cited Count:155 Percentile:78.93(Biochemistry & Molecular Biology)no abstracts in English
Sarai, Akinori*; Siebers, J. G.*; Selvaraj, S.*; Gromiha, M. M.*; Kono, Hidetoshi
Journal of Bioinformatics and Computational Biology, 3(1), p.169 - 183, 2005/02
no abstracts in English
Yonetani, Yoshiteru*; Fujii, Satoshi*; Sarai, Akinori*; Kono, Hidetoshi; Go, Nobuhiro
no journal, ,
DNA conformation and deformability depend on sequence composition. For example, it is known that a sequence containing contiguous AT steps is more deformable. The variability in DNA conformation and deformability has been believed to be one of important factors in specific recognition of DNA sequence by regulatory proteins. Several factors can be considered to yield the difference in DNA conformation and deformability among distinct DNA sequences. One is the mechanical stiffness inherent in the DNA itself, which is originated from base-pairing hydrogen interactions and base-stacking interactions. Another important factor affecting the DNA conformation and deformability is the hydration. The hydration effect is inevitable when discussing the structural properties of DNA, since DNA is physiologically surrounded by water molecules, and the structure cannot be maintained without water. A characteristic hydration pattern observed for DNA is the spine of water, which is composed of the highly ordered water molecules aligned along the floor of the minor groove. Such a hydration pattern as well as the conformation of DNA is dependent on the sequence. Consequently, the following question arises: How and to what extent does the hydration affect the sequence-dependent deformability of DNA ? To address this question, a comparative study of various DNA sequences is required. Recently, the sequence dependence of DNA conformation and deformability has been studied by MD simulations, where all possible 136 patterns for tetramer were examined. In the present work, we report further analysis of these 136 MD trajectories focused on the hydration water. We compared the both properties of DNA hydration and deformation, and found a possible relationship between them.
Yonetani, Yoshiteru*; Fujii, Satoshi*; Sarai, Akinori*; Kono, Hidetoshi; Go, Nobuhiro
no journal, ,
no abstracts in English
Yonetani, Yoshiteru*; Fujii, Satoshi*; Sarai, Akinori*; Kono, Hidetoshi; Go, Nobuhiro
no journal, ,
no abstracts in English
Yonetani, Yoshiteru*; Fujii, Satoshi*; Sarai, Akinori*; Kono, Hidetoshi; Go, Nobuhiro
no journal, ,
no abstracts in English
Yonetani, Yoshiteru; Fujii, Satoshi*; Sarai, Akinori*; Kono, Hidetoshi; Go, Nobuhiro
no journal, ,
no abstracts in English
Yonetani, Yoshiteru*; Fujii, Satoshi*; Sarai, Akinori*; Kono, Hidetoshi; Go, Nobuhiro
no journal, ,
Yonetani, Yoshiteru; Fujii, Satoshi*; Sarai, Akinori*; Kono, Hidetoshi; Go, Nobuhiro*
no journal, ,
no abstracts in English
Fujii, Satoshi*; Kono, Hidetoshi; Go, Nobuhiro*; Sarai, Akinori*
no journal, ,
Yamasaki, Satoshi*; Fukui, Kazuhiko*; Kono, Hidetoshi; Shimizu, Kentaro*; Sarai, Akinori*; Terada, Toru*
no journal, ,