Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Uehara, Akihiro*; Shuhui, X.*; Sato, Ryotaro*; Matsumura, Daiju; Tsuji, Takuya; Yakumaru, Haruko*; Shiro, Ayumi*; Saito, Hiroyuki*; Tanaka, Izumi*; Ishihara, Hiroshi*; et al.
Advances in X-Ray Chemical Analysis, Japan, 53, p.223 - 229, 2022/03
no abstracts in English
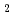 Te
Te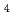 /Bi
/Bi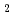 Te
Te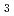 down to 6 K
down to 6 KFukasawa, Takuro*; Kusaka, Shotaro*; Sumida, Kazuki; Hashizume, Mizuki*; Ichinokura, Satoru*; Takeda, Yukiharu; Ideta, Shinichiro*; Tanaka, Kiyohisa*; Shimizu, Ryota*; Hitosugi, Taro*; et al.
Physical Review B, 103(20), p.205405_1 - 205405_6, 2021/05
Times Cited Count:10 Percentile:54.50(Materials Science, Multidisciplinary)Kuroda, Kenta*; Arai, Yosuke*; Rezaei, N.*; Kunisada, So*; Sakuragi, Shunsuke*; Alaei, M.*; Kinoshita, Yuto*; Bareille, C.*; Noguchi, Ryo*; Nakayama, Mitsuhiro*; et al.
Nature Communications (Internet), 11, p.2888_1 - 2888_9, 2020/06
Times Cited Count:36 Percentile:80.96(Multidisciplinary Sciences)Tashiro, Koji*; Kusaka, Katsuhiro*; Hosoya, Takaaki*; Ohara, Takashi; Hanesaka, Makoto*; Yoshizawa, Yoshinori*; Yamamoto, Hiroko*; Niimura, Nobuo*; Tanaka, Ichiro*; Kurihara, Kazuo*; et al.
Macromolecules, 51(11), p.3911 - 3922, 2018/06
Times Cited Count:6 Percentile:18.40(Polymer Science)Hirose, Kentaro; Nishio, Katsuhisa; Tanaka, Shoya*; L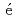 guillon, R.*; Makii, Hiroyuki; Nishinaka, Ichiro*; Orlandi, R.; Tsukada, Kazuaki; Smallcombe, J.*; Vermeulen, M. J.; et al.
guillon, R.*; Makii, Hiroyuki; Nishinaka, Ichiro*; Orlandi, R.; Tsukada, Kazuaki; Smallcombe, J.*; Vermeulen, M. J.; et al.
Physical Review Letters, 119(22), p.222501_1 - 222501_6, 2017/12
Times Cited Count:59 Percentile:91.35(Physics, Multidisciplinary)Fission-fragment mass distributions were measured for 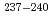 U,
U, 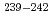 Np and
Np and 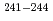 Pu populated in the excitation-energy range from 10 to 60 MeV by multi-nucleon transfer channels in the reaction
Pu populated in the excitation-energy range from 10 to 60 MeV by multi-nucleon transfer channels in the reaction 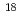 O +
O + 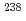 U at the JAEA tandem facility. Among them, the data for
U at the JAEA tandem facility. Among them, the data for 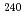 U and
U and 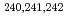 Np were observed for the first time. It was found that the mass distributions for all the studied nuclides maintain a double-humped shape up to the highest measured energy in contrast to expectations of predominantly symmetric fission due to the washing out of nuclear shell effects. From a comparison with the dynamical calculation based on the fluctuation-dissipation model, this behavior of the mass distributions was unambiguously attributed to the effect of multi-chance fission.
Np were observed for the first time. It was found that the mass distributions for all the studied nuclides maintain a double-humped shape up to the highest measured energy in contrast to expectations of predominantly symmetric fission due to the washing out of nuclear shell effects. From a comparison with the dynamical calculation based on the fluctuation-dissipation model, this behavior of the mass distributions was unambiguously attributed to the effect of multi-chance fission.
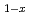 Mn
Mn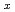 granular thin films
granular thin filmsWakabayashi, Yuki*; Akiyama, Ryota*; Takeda, Yukiharu; Horio, Masafumi*; Shibata, Goro*; Sakamoto, Shoya*; Ban, Yoshisuke*; Saito, Yuji; Yamagami, Hiroshi; Fujimori, Atsushi*; et al.
Physical Review B, 95(1), p.014417_1 - 014417_6, 2017/01
Times Cited Count:11 Percentile:44.75(Materials Science, Multidisciplinary)Tsukuda, Satoshi*; Takahashi, Ryota*; Seki, Shuhei*; Sugimoto, Masaki; Idesaki, Akira; Yoshikawa, Masahito; Tanaka, Shunichiro*
Radiation Physics and Chemistry, 118, p.16 - 20, 2016/01
Times Cited Count:1 Percentile:9.49(Chemistry, Physical)Polyvinylpyrrolidone (PVP)-Pt nanoparticles (NPs) hybrid nanowires were fabricated by high energy ion beam irradiation to PVP thin films including H PtCl
PtCl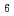 . Single ion hitting caused crosslinking reactions of PVP and reduction of Pt ions within local cylindrical area along an ion trajectory (ion track); therefore, the PVP nanowires including Pt NPs were formed and isolated on Si substrate after wet-development procedure. The number of Pt NPs was easily controlled by the mixed ratio of PVP and H
. Single ion hitting caused crosslinking reactions of PVP and reduction of Pt ions within local cylindrical area along an ion trajectory (ion track); therefore, the PVP nanowires including Pt NPs were formed and isolated on Si substrate after wet-development procedure. The number of Pt NPs was easily controlled by the mixed ratio of PVP and H PtCl
PtCl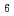 . However, increasing the amount of H
. However, increasing the amount of H PtCl
PtCl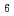 led to decreasing the radial size and separation of the hybrid nanowires during the wet-development. Additional electron beam irradiation after ion beam improved separation of the nanowires and controlled radial sizes due to an increase in the density of crosslinking points inner the nanowires.
led to decreasing the radial size and separation of the hybrid nanowires during the wet-development. Additional electron beam irradiation after ion beam improved separation of the nanowires and controlled radial sizes due to an increase in the density of crosslinking points inner the nanowires.
Unno, Masayoshi*; Ishikawa, Kumiko*; Kusaka, Katsuhiro*; Tamada, Taro; Hagiwara, Yoshinori*; Sugishima, Masakazu*; Wada, Kei*; Yamada, Taro*; Tomoyori, Katsuaki; Hosoya, Takaaki*; et al.
Journal of the American Chemical Society, 137(16), p.5452 - 5460, 2015/04
Times Cited Count:29 Percentile:62.63(Chemistry, Multidisciplinary)Phycocyanobilin, a light-harvesting and photoreceptor pigment in higher plants, algae, and cyanobacteria, is synthesized from biliverdin IX (BV) by phycocyanobilin:ferredoxin oxidoreductase (PcyA) via two steps of two-proton-coupled two-electron reduction. We determined the neutron structure of PcyA from cyanobacteria complexed with BV, revealing the exact location of the hydrogen atoms involved in catalysis. Notably, approximately half of the BV bound to PcyA was BVH
(BV) by phycocyanobilin:ferredoxin oxidoreductase (PcyA) via two steps of two-proton-coupled two-electron reduction. We determined the neutron structure of PcyA from cyanobacteria complexed with BV, revealing the exact location of the hydrogen atoms involved in catalysis. Notably, approximately half of the BV bound to PcyA was BVH , a state in which all four pyrrole nitrogen atoms were protonated. The protonation states of BV complemented the protonation of adjacent Asp105. The "axial "water molecule that interacts with the neutral pyrrole nitrogen of the A-ring was identified. His88 N
, a state in which all four pyrrole nitrogen atoms were protonated. The protonation states of BV complemented the protonation of adjacent Asp105. The "axial "water molecule that interacts with the neutral pyrrole nitrogen of the A-ring was identified. His88 N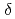 was protonated to form a hydrogen bond with the lactam O atom of the BV A-ring. His88 and His74 were linked by hydrogen bonds via H
was protonated to form a hydrogen bond with the lactam O atom of the BV A-ring. His88 and His74 were linked by hydrogen bonds via H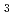 O
O . These results imply that Asp105, His88, and the axial water molecule contribute to proton transfer during PcyA catalysis.
. These results imply that Asp105, His88, and the axial water molecule contribute to proton transfer during PcyA catalysis.
Tashiro, Koji*; Hanesaka, Makoto*; Yamamoto, Hiroko*; Wasanasuk, K.*; Jayaratri, P.*; Yoshizawa, Yoshinori*; Tanaka, Ichiro*; Niimura, Nobuo*; Kusaka, Katsuhiro*; Hosoya, Takaaki*; et al.
Kobunshi Rombunshu, 71(11), p.508 - 526, 2014/11
Times Cited Count:6 Percentile:20.53(Polymer Science)The crystal structure analysis of various polymer substances has been reviewed on the basis of wide-angle high-energy X-ray and neutron diffraction data. The progress in structural analytical techniques of polymer crystals have been reviewed at first. The structural models proposed so far were reinvestigated and new models have been proposed for various kinds of polymer crystals including polyethylene, poly(vinyl alcohol), poly(lactic acid) and its stereocomplex etc. The hydrogen atomic positions were also clarified by the quantitative analysis of wide-angle neutron diffraction data, from which the physical properties of polymer crystals have been evaluated theoretically. The bonded electron density distribution has been estimated for a polydiacetylene single crystal on the basis of the so-called X-N method or by the combination of structural information derived from X-ray and neutron diffraction data analysis. Some comments have been added about future developments in the field of structure-property relationship determination.
Tanaka, Ryota*; Ishihara, Ryo*; Miyoshi, Kazuyoshi*; Umeno, Daisuke*; Saito, Kyoichi*; Asai, Shiho; Yamada, Shinsuke*; Hirota, Hideyuki*
Separation Science and Technology, 49(1), p.154 - 159, 2014/01
Times Cited Count:4 Percentile:15.89(Chemistry, Multidisciplinary)Tokuhisa, Atsushi*; Arai, Junya*; Jochi, Yasumasa*; Ono, Yoshiyuki*; Kameyama, Toyohisa*; Yamamoto, Keiji*; Hatanaka, Masayuki*; Gerofi, B.*; Shimada, Akio*; Kurokawa, Motoyoshi*; et al.
Journal of Synchrotron Radiation, 20(6), p.899 - 904, 2013/11
Times Cited Count:5 Percentile:27.82(Instruments & Instrumentation) -thrombin-bivalirubin complex at pD 5.0; Protonation states and hydration structure of the enzyme-product complex
-thrombin-bivalirubin complex at pD 5.0; Protonation states and hydration structure of the enzyme-product complexYamada, Taro*; Kurihara, Kazuo; Onishi, Yuki*; Tamada, Taro; Tomoyori, Katsuaki; Masumi, Kenji*; Tanaka, Ichiro*; Kuroki, Ryota; Niimura, Nobuo*
Biochimica et Biophysica Acta; Proteins and Proteomics, 1834(8), p.1532 - 1538, 2013/08
Times Cited Count:20 Percentile:52.35(Biochemistry & Molecular Biology)The protonation states and hydration structures of the  -thrombin-bivalirubin complex were studied by joint XN refinement of the single crystal X-ray and neutron diffraction data at resolutions of 1.6 and 2.8
-thrombin-bivalirubin complex were studied by joint XN refinement of the single crystal X-ray and neutron diffraction data at resolutions of 1.6 and 2.8  , respectively. The atomic distances were estimated by carrying out X-ray crystallographic analysis at 1.25
, respectively. The atomic distances were estimated by carrying out X-ray crystallographic analysis at 1.25  resolution. The complex represents a model of the enzyme-product (EP) complex of
resolution. The complex represents a model of the enzyme-product (EP) complex of  -thrombin. The neutron scattering length maps around the active site suggest that the side chain of H57/H was deuterated. The joint XN refinement showed that occupancies for D
-thrombin. The neutron scattering length maps around the active site suggest that the side chain of H57/H was deuterated. The joint XN refinement showed that occupancies for D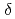 1 and D
1 and D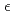 2 of H57/H were 1.0 and 0.7, respectively. However, no significant neutron scattering length density was observed around the hydroxyl oxygen O
2 of H57/H were 1.0 and 0.7, respectively. However, no significant neutron scattering length density was observed around the hydroxyl oxygen O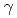 of S195/H, which was close to the carboxylic carbon atom of dFPR-COOH. These observations suggest that the O
of S195/H, which was close to the carboxylic carbon atom of dFPR-COOH. These observations suggest that the O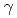 atom of S195/H is deprotonated and maintains its nucleophilicity in the EP complex. In addition to the active site, the hydration structures of the S1 subsite and the Exosite I, which are involved in the recognition of bivalirudin, are presented.
atom of S195/H is deprotonated and maintains its nucleophilicity in the EP complex. In addition to the active site, the hydration structures of the S1 subsite and the Exosite I, which are involved in the recognition of bivalirudin, are presented.
Honda, Ryotaro; Callier, S.*; Hasegawa, Shoichi; Ieiri, Masaharu*; Matsumoto, Yuki*; Miwa, Koji*; Nakamura, Isamu*; Raux, L.*; De La Taille, C.*; Tanaka, Manobu*; et al.
Nuclear Instruments and Methods in Physics Research A, 695, p.206 - 209, 2012/12
Times Cited Count:3 Percentile:23.63(Instruments & Instrumentation)For the J-PARC E40 experiment which aims to measure differential cross-sections of Sp scatterings, a system to detect scattered proton from Sp scatterings is under development. The detection system consists of scintillation fibers with a MPPC readout. A prototype and a readout electronics for MPPC have already been developed. The prototype consisting of a scintillation fiber tracker and a BGO calorimeter was tested with a proton beam of 80 MeV. Energy resolutions of the tracker of 22.0% (s) and the calorimeter of 1.0% (s) were obtained for 1 MeV and 70 MeV energy deposit, respectively. The prototype readout electronics has an ASIC for multichannel operation, EASIROC, and a Silicon TCP (SiTCP) interface to communicate with a DAQ system. Its data transfer rate measured was 14 kHz. Required performances for the prototype system have been achieved except for the energy resolution of the prototype fiber tracker.
Kawamura, Kenji*; Yamada, Taro*; Kurihara, Kazuo; Tamada, Taro; Kuroki, Ryota; Tanaka, Ichiro*; Takahashi, Haruyuki*; Niimura, Nobuo*
Acta Crystallographica Section D, 67(2), p.140 - 148, 2011/02
Times Cited Count:30 Percentile:88.81(Biochemical Research Methods)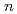 -octylphosphine oxide, and its adsorption/elution of bismuth ions
-octylphosphine oxide, and its adsorption/elution of bismuth ionsTanaka, Ryota*; Ishihara, Ryo*; Miyoshi, Kazuyoshi*; Umeno, Daisuke*; Saito, Kyoichi*; Asai, Shiho; Yamada, Shinsuke*; Hirota, Hideyuki*
Reactive and Functional Polymers, 70(12), p.986 - 990, 2010/12
Times Cited Count:7 Percentile:28.05(Chemistry, Applied) resolution
resolutionYagi, Daichi*; Yamada, Taro*; Kurihara, Kazuo; Onishi, Yuki*; Yamashita, Masahiro*; Tamada, Taro; Tanaka, Ichiro*; Kuroki, Ryota; Niimura, Nobuo*
Acta Crystallographica Section D, 65(9), p.892 - 899, 2009/09
Times Cited Count:17 Percentile:79.69(Biochemical Research Methods)A neutron crystallographic analysis of phosphate-free bovine pancreatic RNase A has been carried out at 1.7  resolution using the BIX-4 single-crystal diffractometer at the JRR-3 reactor of the Japan Atomic Energy Agency. The high resolution structural model allowed us to determine that His12 acts mainly as a general base in the catalytic process of RNase A. Numerous other distinctive structural features such as the hydrogen positions of methyl groups, hydroxyl groups, prolines, asparagines and glutamines were also determined at 1.7
resolution using the BIX-4 single-crystal diffractometer at the JRR-3 reactor of the Japan Atomic Energy Agency. The high resolution structural model allowed us to determine that His12 acts mainly as a general base in the catalytic process of RNase A. Numerous other distinctive structural features such as the hydrogen positions of methyl groups, hydroxyl groups, prolines, asparagines and glutamines were also determined at 1.7  resolution. The protonation and deprotonation states of all of the charged amino-acid residues allowed us to provide a definitive description of the hydrogen-bonding network around the active site and the H atoms of the key His48 residue. Differences in hydrogen-bond strengths for the
resolution. The protonation and deprotonation states of all of the charged amino-acid residues allowed us to provide a definitive description of the hydrogen-bonding network around the active site and the H atoms of the key His48 residue. Differences in hydrogen-bond strengths for the  -helices and
-helices and  -sheets were inferred from determination of the hydrogen-bond lengths and the H/D-exchange ratios of the backbone amide H atoms. The correlation between the B factors and hydrogen-bond lengths of the hydration water molecules was also determined.
-sheets were inferred from determination of the hydrogen-bond lengths and the H/D-exchange ratios of the backbone amide H atoms. The correlation between the B factors and hydrogen-bond lengths of the hydration water molecules was also determined.
Ito, Takachika*; Suzuki, Shoichi*; Kanaji, Sachiko*; Shiraishi, Hiroshi*; Ota, Shoichiro*; Arima, Kazuhiko*; Tanaka, Go*; Tamada, Taro; Honjo, Eijiro*; Garcia, K. C.*; et al.
Journal of Biological Chemistry, 284(36), p.24289 - 24296, 2009/09
Times Cited Count:24 Percentile:45.35(Biochemistry & Molecular Biology)Both IL-4 and IL-13 can bind to the shared receptor composed of the IL-4 receptor  chain and the IL-13 receptor
chain and the IL-13 receptor  -1 chain (IL-13R
-1 chain (IL-13R 1); however, the assembly mechanisms of these ligands to the receptor is different, enabling the principal functions of these ligands to be different. We have previously shown that the N-terminal Ig-like domain in IL-13R
1); however, the assembly mechanisms of these ligands to the receptor is different, enabling the principal functions of these ligands to be different. We have previously shown that the N-terminal Ig-like domain in IL-13R 1, called the D1 domain, is the specific and critical binding unit for IL-13. However, it has still remained obscure which the amino acid has specific binding capacity to IL-13 and why the D1 domain acts as the binding site for IL-13, but not IL-4. To address these questions, in this study, we performed the mutational analyses for the D1 domain, combining the structural data to identify the amino acids critical for binding to IL-13. Mutations of Lys76, Lys77, or Ile78 in c' strand in which the crystal structure showed interact with IL-13 and those of Trp65 and Ala79 adjacent to the interacting site, resulted in significant impairment of IL-13 binding, demonstrating that these amino acids generate the binding site. Furthermore, mutations of Val35, Leu38, or Val42 at N-terminal
1, called the D1 domain, is the specific and critical binding unit for IL-13. However, it has still remained obscure which the amino acid has specific binding capacity to IL-13 and why the D1 domain acts as the binding site for IL-13, but not IL-4. To address these questions, in this study, we performed the mutational analyses for the D1 domain, combining the structural data to identify the amino acids critical for binding to IL-13. Mutations of Lys76, Lys77, or Ile78 in c' strand in which the crystal structure showed interact with IL-13 and those of Trp65 and Ala79 adjacent to the interacting site, resulted in significant impairment of IL-13 binding, demonstrating that these amino acids generate the binding site. Furthermore, mutations of Val35, Leu38, or Val42 at N-terminal  -strand also resulted in loss of IL-13 binding, probably from decrease structural stability. None of the mutations employed here affected IL-4 binding. These results demonstrate that the hydrophobic patch composed of Lys76, Lys77, and Ile78 is the IL-13 recognition site and solidify our understanding that the differential requirements of the D1 domain in IL-13R
-strand also resulted in loss of IL-13 binding, probably from decrease structural stability. None of the mutations employed here affected IL-4 binding. These results demonstrate that the hydrophobic patch composed of Lys76, Lys77, and Ile78 is the IL-13 recognition site and solidify our understanding that the differential requirements of the D1 domain in IL-13R 1 allows the shared receptor to respond differentially to IL-4 and IL-13.
1 allows the shared receptor to respond differentially to IL-4 and IL-13.
Hidaka, Koshi*; Kimura, Toru*; Abdel-Rahman, H. M.*; Nguyen, J.-T.*; McDaniel, K. F.*; Kohlbrenner, W. E.*; Molla, A.*; Adachi, Motoyasu; Tamada, Taro; Kuroki, Ryota; et al.
Journal of Medicinal Chemistry, 52(23), p.7604 - 7617, 2009/07
Times Cited Count:20 Percentile:44.78(Chemistry, Medicinal)A series of HIV protease inhibitor based on the allophenylnorstatine structure with various P2' moieties were synthesized. Among these analogues, we discovered that a small allyl group would maintain potent enzyme inhibitory activity compared to that of the 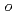 -methylbenzyl moiety in clinical candidate 1 (KNI-764, also known as JE-2147, AG-1776 or SM-319777). Introduction of an anilinic amino group to 2 (KNI-727) improved water-solubility and anti-HIV-1 activity. X-ray crystallographic analysis of 13k (KNI-1689) with a
-methylbenzyl moiety in clinical candidate 1 (KNI-764, also known as JE-2147, AG-1776 or SM-319777). Introduction of an anilinic amino group to 2 (KNI-727) improved water-solubility and anti-HIV-1 activity. X-ray crystallographic analysis of 13k (KNI-1689) with a  -methallyl group at P2' position revealed hydrophobic interactions with Ala28, Ile84, and Ile50' similar to that of 1. The presence of an additional methyl group on the allyl group in compound 13k significantly increased anti-HIV activity over 1, while providing a rational drug design for structural minimization and improving membrane permeability.
-methallyl group at P2' position revealed hydrophobic interactions with Ala28, Ile84, and Ile50' similar to that of 1. The presence of an additional methyl group on the allyl group in compound 13k significantly increased anti-HIV activity over 1, while providing a rational drug design for structural minimization and improving membrane permeability.
 chain and interleukin-13 receptor
chain and interleukin-13 receptor  1 chain by a silkworm-baculovirus system
1 chain by a silkworm-baculovirus systemHonjo, Eijiro; Shoyama, Yoshinari; Tamada, Taro; Shigematsu, Hideki*; Hatanaka, Takaaki*; Kanaji, Sachiko*; Arima, Kazuhiko*; Ito, Yuji*; Izuhara, Kenji*; Kuroki, Ryota
Protein Expression and Purification, 60(1), p.25 - 30, 2008/07
Times Cited Count:13 Percentile:32.55(Biochemical Research Methods)The receptor binding to Interleukin (IL)-13 is composed of the IL-13 receptor  1 chain (IL-13R
1 chain (IL-13R  1) and the IL-4 receptor
1) and the IL-4 receptor  chain (IL-4R
chain (IL-4R  ). In order to investigate the interaction of IL-13 with IL-13R
). In order to investigate the interaction of IL-13 with IL-13R  1 and IL-4R
1 and IL-4R  , the DNA fragments coding the extracellular regions of human IL-13R
, the DNA fragments coding the extracellular regions of human IL-13R  1 and the IL-4R
1 and the IL-4R  were fused with mouse Fc and expressed by a silkworm-baculovirus system. The expressed receptors were successfully purified by affinity chromatography using protein A, and the Fc region was removed by thrombin digestion. Size exclusion chromatography and SPR analysis revealed that mixture of IL-13 and IL-13R
were fused with mouse Fc and expressed by a silkworm-baculovirus system. The expressed receptors were successfully purified by affinity chromatography using protein A, and the Fc region was removed by thrombin digestion. Size exclusion chromatography and SPR analysis revealed that mixture of IL-13 and IL-13R 1 showed predominant affinity to IL-4R
1 showed predominant affinity to IL-4R , although neither detectable affinity of IL-13 nor IL-13R
, although neither detectable affinity of IL-13 nor IL-13R 1 was observed against IL-4R
1 was observed against IL-4R . Combining these data with the moderate affinity of IL-13 to IL-13R
. Combining these data with the moderate affinity of IL-13 to IL-13R 1, this indicates that IL-13 first binds to IL-13R
1, this indicates that IL-13 first binds to IL-13R 1 and recruits consequently to IL-4R.
1 and recruits consequently to IL-4R.
Ishikawa, Takuya*; Chatake, Toshiyuki*; Onishi, Yuki*; Tanaka, Ichiro*; Kurihara, Kazuo; Kuroki, Ryota; Niimura, Nobuo*
Chemical Physics, 345(2-3), p.152 - 158, 2008/04
Times Cited Count:14 Percentile:43.46(Chemistry, Physical)