Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Tanaka, Kazuya; Yamaji, Keiko*; Masuya, Hayato*; Tomita, Jumpei; Ozawa, Mayumi*; Yamasaki, Shinya*; Tokunaga, Kohei; Fukuyama, Kenjin*; Ohara, Yoshiyuki*; Maamoun, I.*; et al.
Chemosphere, 355, p.141837_1 - 141837_11, 2024/05
In this study, biogenic Mn(IV) oxide was applied to remove Ra from mine water collected from a U mill tailings pond in the Ningyo-toge center. Just 7.6 mg of biogenic Mn(IV) oxide removed more than 98% of the 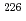 Ra from 3 L of mine water, corresponding to a distribution coefficient of 10
Ra from 3 L of mine water, corresponding to a distribution coefficient of 10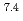 mL/g for Ra at pH 7. The obtained value was convincingly high for practical application of biogenic Mn(IV) oxide in water treatment.
mL/g for Ra at pH 7. The obtained value was convincingly high for practical application of biogenic Mn(IV) oxide in water treatment.
Ogura, Koya*; Hosoda, Masahiro*; Tamakuma, Yuki*; Suzuki, Takahito*; Yamada, Ryohei; Negemi, Ryoju*; Tsujiguchi, Takakiyo*; Yamaguchi, Masaru*; Shiroma, Yoshitaka*; Iwaoka, Kazuki*; et al.
International Journal of Environmental Research and Public Health, 18(3), p.978_1 - 978_16, 2021/02
Times Cited Count:7 Percentile:68.83(Environmental Sciences)Yamasaki, Chisato*; Murakami, Katsuhiko*; Fujii, Yasuyuki*; Sato, Yoshiharu*; Harada, Erimi*; Takeda, Junichi*; Taniya, Takayuki*; Sakate, Ryuichi*; Kikugawa, Shingo*; Shimada, Makoto*; et al.
Nucleic Acids Research, 36(Database), p.D793 - D799, 2008/01
Times Cited Count:51 Percentile:71.25(Biochemistry & Molecular Biology)Here we report the new features and improvements in our latest release of the H-Invitational Database, a comprehensive annotation resource for human genes and transcripts. H-InvDB, originally developed as an integrated database of the human transcriptome based on extensive annotation of large sets of fulllength cDNA (FLcDNA) clones, now provides annotation for 120 558 human mRNAs extracted from the International Nucleotide Sequence Databases (INSD), in addition to 54 978 human FLcDNAs, in the latest release H-InvDB. We mapped those human transcripts onto the human genome sequences (NCBI build 36.1) and determined 34 699 human gene clusters, which could define 34 057 protein-coding and 642 non-protein-coding loci; 858 transcribed loci overlapped with predicted pseudogenes.
Yamaguchi, Tetsuji; Sakamoto, Yoshifumi; Akai, Masanobu; Takazawa, Mayumi; Iida, Yoshihisa; Tanaka, Tadao; Nakayama, Shinichi
Physics and Chemistry of the Earth, 32(1-7), p.298 - 310, 2007/00
Times Cited Count:43 Percentile:72.99(Geosciences, Multidisciplinary)Dissolution rate of montmorillonite, diffusivity of hydroxide ion and permeability coefficient in compacted sand-bentonite mixtures were experimentally determined and formulated. A coupled mass-transport/chemical-reaction code was developed to predict variation in permeability of engineered bentonite barrier with alkaline fluid by using the formulae.
Tanaka, Tadao; Sakamoto, Yoshifumi; Yamaguchi, Tetsuji; Takazawa, Mayumi; Akai, Masanobu; Negishi, Kumi; Iida, Yoshihisa; Nakayama, Shinichi
JAERI-Conf 2005-007, p.105 - 110, 2005/08
Highly alkaline environments induced by cementitious materials in radioactive waste repositories are likely to dissolve and to alter montmorillonite, the main constituent of bentonite buffer materials. For the prediction of the long-term variations in permeability of compacted sand-bentonite mixtures, long-term alteration of bentonite should be quantified based on information accumulated by using the compacted or powdered bentonite materials, with batch experiments or column experiments. In this study, we summarize distinctive information obtained from various experimental systems, and propose functional and effective integration of experimental approaches to prediction of bentonite alteration.
Takazawa, Mayumi; Negishi, Kumi; Sakamoto, Yoshifumi; Akai, Masanobu; Yamaguchi, Tetsuji; Iida, Yoshihisa; Tanaka, Tadao; Nakayama, Shinichi
JAERI-Conf 2005-007, p.236 - 241, 2005/08
no abstracts in English
Takazawa, Mayumi; Yamaguchi, Tetsuji; Sakamoto, Yoshifumi; Akai, Masanobu; Tanaka, Tadao; Nakayama, Shinichi
NUMO-TR-04-05, p.A3_59 - A3_62, 2004/10
no abstracts in English
Nishimura, Akihiko; Yamaguchi, Toshihiko; Oka, Kiyoshi; Akatsu, Tomohiro*; Ito, Fuyumi; Yamashita, Takuya
no journal, ,
no abstracts in English
Nishimura, Akihiko; Yamaguchi, Toshihiko; Akatsu, Tomohiro*; Ito, Fuyumi; Oka, Kiyoshi
no journal, ,
no abstracts in English
Nishimura, Akihiko; Akatsu, Tomohiro*; Takenaka, Yusuke*; Yamaguchi, Toshihiko; Ito, Fuyumi; Oka, Kiyoshi; Yamashita, Takuya
no journal, ,
no abstracts in English
Taguchi, Mitsumasa; Iwamatsu, Kazuhiro; Sugo, Yumi; Kurashima, Satoshi; Yamaguchi, Makoto*; Katsumura, Yosuke
no journal, ,
no abstracts in English
Watanabe, Shigeki; Yamada, Keiichi*; Hanaoka, Hirofumi*; Ohshima, Yasuhiro; Takano, Chikako*; Tsukui, Narutaka*; Yamaguchi, Aiko*; Sugo, Yumi; Oku, Hiroyuki*; Ishioka, Noriko
no journal, ,
no abstracts in English
Sugo, Yumi; Ohshima, Yasuhiro; Yamaguchi, Aiko*; Hanaoka, Hirofumi*; Ishioka, Noriko
no journal, ,
no abstracts in English
Hemmi, Ko; Tachibana, Mayumi; Aoyama, Eri; Yamaguchi, Tetsuji; Maeda, Toshikatsu
no journal, ,
no abstracts in English