Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Chiba, Kaori*; Matsui, Takuro*; Chatake, Toshiyuki*; Ohara, Takashi; Tanaka, Ichiro*; Yutani, Katsuhide*; Niimura, Nobuo*
Protein Science, 32(10), p.e4765_1 - e4765_13, 2023/10
Times Cited Count:1 Percentile:5.28(Biochemistry & Molecular Biology)Tashiro, Koji*; Kusaka, Katsuhiro*; Hosoya, Takaaki*; Ohara, Takashi; Hanesaka, Makoto*; Yoshizawa, Yoshinori*; Yamamoto, Hiroko*; Niimura, Nobuo*; Tanaka, Ichiro*; Kurihara, Kazuo*; et al.
Macromolecules, 51(11), p.3911 - 3922, 2018/06
Times Cited Count:8 Percentile:23.37(Polymer Science) resolution
resolutionChatake, Toshiyuki*; Kurihara, Kazuo; Tanaka, Ichiro*; Tsyba, I.*; Bau, R.*; Jenney, F. E. Jr.*; Adams, M. W. W.*; Niimura, Nobuo
Acta Crystallographica Section D, 60(8), p.1364 - 1373, 2004/08
Times Cited Count:34 Percentile:88.25(Biochemical Research Methods)A neutron diffraction study has been carried out at 1.6  resolution on a mutant rubredoxin from
resolution on a mutant rubredoxin from 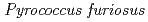 using the BIX-3 single-crystal diffractometer at the JRR-3 reactor of JAERI. In order to study the unusual thermostability of rubredoxin from
using the BIX-3 single-crystal diffractometer at the JRR-3 reactor of JAERI. In order to study the unusual thermostability of rubredoxin from 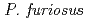 , the hydrogen-bonding patterns were compared between the native and a 'triple-mutant' variant where three residues were changed so that they are identical to those in a mesophilic rubredoxin. In the present study, some minor changes were found between the wild-type and mutant proteins in the hydrogen-bonding patterns of the Trp3/Tyr3 region. The H/D-exchange ratios in the protein were also studied. The results suggest that the backbone amide bonds near the four Cys residues of the FeS
, the hydrogen-bonding patterns were compared between the native and a 'triple-mutant' variant where three residues were changed so that they are identical to those in a mesophilic rubredoxin. In the present study, some minor changes were found between the wild-type and mutant proteins in the hydrogen-bonding patterns of the Trp3/Tyr3 region. The H/D-exchange ratios in the protein were also studied. The results suggest that the backbone amide bonds near the four Cys residues of the FeS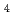 redox center are most resistant to H/D exchange. In addition, the 1.6
redox center are most resistant to H/D exchange. In addition, the 1.6  resolution of the present neutron structure determination has revealed a more detailed picture than previously available of some portions of the water structure, including ordered and disordered O-D bonds.
resolution of the present neutron structure determination has revealed a more detailed picture than previously available of some portions of the water structure, including ordered and disordered O-D bonds.
Kurihara, Kazuo; Tanaka, Ichiro; Adams, M. W. W.*; Jenney, F. E. Jr.*; Moiseeva, N.*; Bau, R.*; Niimura, Nobuo
Journal of the Physical Society of Japan, Vol.70, Supplement A, p.400 - 402, 2001/05
With the new single-crystal diffractometer BIX-3 at the JRR-3M reactor of JAERI, a single-crystal neutron diffraction analysis of the structure of the small protein rubredoxin from the hyperthermophile Pyrococcus furiosus is currently under way. Data were collected at room temperature up to a resolution of 1.5  intervals in
intervals in 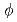 and exposure times ranging from 60 to 77 minutes per frame. The completeness factor of the 1.5-
and exposure times ranging from 60 to 77 minutes per frame. The completeness factor of the 1.5- . Included in the refinement are 301 hydrogen atoms and 40 deuterium atoms, and 29 water molecules were also identified. In the present model, the current value for R and R
. Included in the refinement are 301 hydrogen atoms and 40 deuterium atoms, and 29 water molecules were also identified. In the present model, the current value for R and R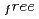 are 24.0
are 24.0 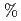 and 26.3
and 26.3 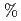 , respectively.
, respectively.
Kurihara, Kazuo*; Kono, Fumiaki*; Shimizu, Rumi*; Tamura, Itaru; Tamada, Taro*
no journal, ,
no abstracts in English
Kurihara, Kazuo*; Kono, Fumiaki*; Shimizu, Rumi*; Tamura, Itaru; Tamada, Taro*
no journal, ,
no abstracts in English
Kurihara, Kazuo*; Kono, Fumiaki*; Shimizu, Rumi*; Tamura, Itaru; Tamada, Taro*
no journal, ,
no abstracts in English
Kurihara, Kazuo*; Hirano, Yu*; Hiromoto, Takeshi*; Tamura, Itaru; Tamada, Taro*
no journal, ,
no abstracts in English
Kurihara, Kazuo*; Kono, Fumiaki*; Shimizu, Rumi*; Tamura, Itaru; Tamada, Taro*
no journal, ,
no abstracts in English
Kurihara, Kazuo*; Kono, Fumiaki*; Shimizu, Rumi*; Tamura, Itaru; Ohara, Takashi; Chatake, Toshiyuki*; Inoue, Rintaro*; Hirano, Yu*; Tamada, Taro*
no journal, ,