Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Nagamatsu, Aiko*; Casolino, M.*; Larsson, O.*; Ito, Tsuyoshi*; Yasuda, Nakahiro*; Kitajo, Keiichi*; Shimada, Ken*; Takeda, Kazuo*; Tsuda, Shuichi; Sato, Tatsuhiko
Physics Procedia, 80, p.25 - 35, 2015/12
Times Cited Count:12 Percentile:95.25(Physics, Applied)JAXA has been investigated the effect of the shielding material for dosimeters in the ISS Russian segment under the ALTCRISS project. The PADLES package is a passive dosimeters which usually consists of two types of passive and integrating dosimeters, a CR-39 plastic nuclear track detector and a thermoluminescence dosimeter. In the ALTCRISS experiments, both tissue-equivalent material CR-39 PNTDs and NAN-JAERI were enclosed in the PADLES packages as radiators for CR-39 to precisely measure a personal exposed doses. We compared the doses obtained by PADLESs with or without the PEs, and with or difference of the two tissue-equivalent materials. The results of space radiation measurements of the ALTCRISS project Phase 1 to 4 using the PADLES system will be reported in this time: Phase 1 experiments was from December 2005 to April 2006, Phase 2 was from April to September 2006, phase 3 was from Sep 2006 to April 2007, and Phase 4 was conducted from May to October 2007.
 hypersensitive and hypermutable in response to UV-B radiation
hypersensitive and hypermutable in response to UV-B radiationYoshihara, Ryohei; Nakane, Chiyoko*; Sato, Ryohei*; Yasuda, Ai*; Takimoto, Koichi*
Genes and Environment, 30(2), p.53 - 61, 2008/05
We generated a CPD photolyase knock-down in 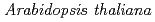 by RNAi to investigate the role of CPD photorepair for protection of plants from solar UV. These knock-down lines exhibited hypersensitivity to UV-B and an increased occurrence of mutation induced by UV-B radiation compared with
by RNAi to investigate the role of CPD photorepair for protection of plants from solar UV. These knock-down lines exhibited hypersensitivity to UV-B and an increased occurrence of mutation induced by UV-B radiation compared with 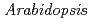 proficient in CPD photolyase. These results indicate that CPD photoreactivation plays an important role for UV resistance of
proficient in CPD photolyase. These results indicate that CPD photoreactivation plays an important role for UV resistance of 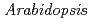 and suppression of UV-induced mutagenesis.
and suppression of UV-induced mutagenesis.
Yamasaki, Chisato*; Murakami, Katsuhiko*; Fujii, Yasuyuki*; Sato, Yoshiharu*; Harada, Erimi*; Takeda, Junichi*; Taniya, Takayuki*; Sakate, Ryuichi*; Kikugawa, Shingo*; Shimada, Makoto*; et al.
Nucleic Acids Research, 36(Database), p.D793 - D799, 2008/01
Times Cited Count:51 Percentile:71.25(Biochemistry & Molecular Biology)Here we report the new features and improvements in our latest release of the H-Invitational Database, a comprehensive annotation resource for human genes and transcripts. H-InvDB, originally developed as an integrated database of the human transcriptome based on extensive annotation of large sets of fulllength cDNA (FLcDNA) clones, now provides annotation for 120 558 human mRNAs extracted from the International Nucleotide Sequence Databases (INSD), in addition to 54 978 human FLcDNAs, in the latest release H-InvDB. We mapped those human transcripts onto the human genome sequences (NCBI build 36.1) and determined 34 699 human gene clusters, which could define 34 057 protein-coding and 642 non-protein-coding loci; 858 transcribed loci overlapped with predicted pseudogenes.
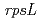 gene in plant
gene in plantYoshihara, Ryohei; Yasuda, Ai*; Sato, Ryohei*; Hase, Yoshihiro; Narumi, Issei; Takimoto, Koichi*
no journal, ,
no abstracts in English
Sato, Ryohei*; Yasuda, Ai*; Yoshihara, Ryohei; Takimoto, Koichi*
no journal, ,
no abstracts in English