Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
 MnSi
MnSiShigeta, Iduru*; Oku, Shuta*; Kubota, Takahide*; Kimura, Shojiro*; Seki, Takeshi*; Shinozaki, Bunju*; Awaji, Satoshi*; Takanashi, Koki; Hiroi, Masahiko*
AIP Advances (Internet), 13(2), p.025116_1 - 025116_5, 2023/02
Times Cited Count:1 Percentile:13.37(Nanoscience & Nanotechnology)Nakayama, Shigeru*; Kawaguchi, Koichi; Segawa, Tomoomi; Yamada, Yoshikazu
Proceedings of 19th International Symposium on Artificial Life and Robotics (AROB 2014) (CD-ROM), p.246 - 249, 2014/01
In the nuclear fuel fabrication process, moisture content is very important parameter because of criticality safety control. Considering future commercial nuclear fuel fabrication plant, rapid and durable moisture sensor is required. We have developed an open-ended semi-coaxial microwave sensor to measure moisture involved in a substance. This sensor has no semiconductor tips, so it can be used in strong radiation field. In this paper, we carry out a preliminary experiment for measuring moisture of MOX(UO +PuO
+PuO ) in granulation process, in which water is added as a binder. In our preliminary experiment, to simulate granulated MOX powder, granulated tungsten trioxide powder, which has similar dielectric constant to MOX and has voids to hold water inside, was used. The principle of microwave measurement of moisture is as follows. When the tungsten trioxide contained in a pyrex beaker is placed at the open end of the cavity, the resonant frequency is shifted by a variation in the end of capacitance which results from the difference in the dielectric constant of tungsten trioxide from that of air. Furthermore, the peak value of the resonance curve is attenuated by the absorption of microwave in the tungsten trioxide. Therefore, the moisture content of tungsten trioxide can be estimated by measuring either the frequency shift or attenuation. They are measured using a tracking generator and a spectrum analyzer. In our presentation, we will show the experimental results in detail.
) in granulation process, in which water is added as a binder. In our preliminary experiment, to simulate granulated MOX powder, granulated tungsten trioxide powder, which has similar dielectric constant to MOX and has voids to hold water inside, was used. The principle of microwave measurement of moisture is as follows. When the tungsten trioxide contained in a pyrex beaker is placed at the open end of the cavity, the resonant frequency is shifted by a variation in the end of capacitance which results from the difference in the dielectric constant of tungsten trioxide from that of air. Furthermore, the peak value of the resonance curve is attenuated by the absorption of microwave in the tungsten trioxide. Therefore, the moisture content of tungsten trioxide can be estimated by measuring either the frequency shift or attenuation. They are measured using a tracking generator and a spectrum analyzer. In our presentation, we will show the experimental results in detail.
 sp. nucleoside diphosphate kinase
sp. nucleoside diphosphate kinaseArai, Shigeki; Yonezawa, Yasushi; Okazaki, Nobuo; Matsumoto, Fumiko; Tamada, Taro; Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Blaber, M.; Tokunaga, Masao*; Kuroki, Ryota
Protein Science, 21(4), p.498 - 510, 2012/04
Times Cited Count:15 Percentile:34.92(Biochemistry & Molecular Biology)In order to clarify the oligomer state of nucleoside diphosphate kinase (NDK) from moderately halophilic  sp. 593 (HaNDK), the crystal structure of HaNDK was determined by X-ray crystallography. The crystal structures of the wild-type HaNDK and the mutant HaNDK (E134A) showed a dimer and a tetramer, respectively. The higher ordered association of proteins usually contributes to an increase in thermal stability and substrate affinity. The change in the assembly form by a minimum mutation may be an effective way for NDK to acquire molecular characteristics suited to various circumstances.
sp. 593 (HaNDK), the crystal structure of HaNDK was determined by X-ray crystallography. The crystal structures of the wild-type HaNDK and the mutant HaNDK (E134A) showed a dimer and a tetramer, respectively. The higher ordered association of proteins usually contributes to an increase in thermal stability and substrate affinity. The change in the assembly form by a minimum mutation may be an effective way for NDK to acquire molecular characteristics suited to various circumstances.
Nakata, Toshiya; Komazaki, Shinichi*; Kono, Yutaka*; Tanigawa, Hiroyasu
Metallurgical Journal, LXIII(Sp.), p.146 - 150, 2010/08
The small punch (SP) test is one method of small specimen test technology. We constructed a model for finite element analysis (FEA) using the Ramberg-Osgood equation to try to estimate tensile properties from the SP test. The results showed that the SP curve obtained from previous experiments could be reproduced by FEA to the point where the necking behavior of SP specimens became prominent. It was found that the relationship between the Mises equivalent stress at the point giving the largest equivalent plastic strain, and the total strain based on the Ramberg-Osgood equation, in the SP specimens, agreed well with the true stress-true strain curve plotted from tensile test measurements, and could thus be used to evaluate the 0.2% proof stress, tensile strength and uniform elongation.
 DSM3043; Both residues 134 and 136 are critical for the tetramer assembly
DSM3043; Both residues 134 and 136 are critical for the tetramer assemblyTokunaga, Hiroko*; Izutsu, Kenichi*; Arai, Shigeki; Yonezawa, Yasushi; Kuroki, Ryota; Arakawa, Tsutomu*; Tokunaga, Masao*
Enzyme and Microbial Technology, 46(2), p.129 - 135, 2010/02
Times Cited Count:6 Percentile:20.05(Biotechnology & Applied Microbiology)Both wild-type nucleoside diphosphate kinase from moderately halophilc  (CsNDK (GNE), GNE represents Gly134-Asn135-Glu136) and mutant CsNDK (ANE), both of which have a neutral amino acid at residue 134, were found to form a dimer. These constructs contain Glu136, which may also cause steric barrier and charge repulsion. A double mutant, CsNDK (ANT), having Thr at 136 resulted in stable tetrameric assembly, supporting the above notion. A mutant CsNDK (GNT) reverted, however, to a dimer again, indicating that the introduced Ala residue at 134th in the double mutant generated a hydrophobic cluster consisting of the Ala residues and thereby stabilized dimer-dimer association of CsNDK assembly, while Gly destabilized it due to the loss of this cluster. Based on these observations, it is evident that both residues 134 and 136 contribute to the subunit assembly of CsNDK.
(CsNDK (GNE), GNE represents Gly134-Asn135-Glu136) and mutant CsNDK (ANE), both of which have a neutral amino acid at residue 134, were found to form a dimer. These constructs contain Glu136, which may also cause steric barrier and charge repulsion. A double mutant, CsNDK (ANT), having Thr at 136 resulted in stable tetrameric assembly, supporting the above notion. A mutant CsNDK (GNT) reverted, however, to a dimer again, indicating that the introduced Ala residue at 134th in the double mutant generated a hydrophobic cluster consisting of the Ala residues and thereby stabilized dimer-dimer association of CsNDK assembly, while Gly destabilized it due to the loss of this cluster. Based on these observations, it is evident that both residues 134 and 136 contribute to the subunit assembly of CsNDK.
Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Arisaka, Fimio*; Arai, Shigeki; Kuroki, Ryota; Arakawa, Tsutomu*; Tokunaga, Masao*
FEBS Letters, 582(7), p.1049 - 1054, 2008/04
Times Cited Count:17 Percentile:39.41(Biochemistry & Molecular Biology) nucleoside diphosphate kinase (HaNDK) forms a dimeric assembly and
nucleoside diphosphate kinase (HaNDK) forms a dimeric assembly and  NDK (PaNDK) forms a tetrameric assembly. The mutation of Glu134 to Ala in HaNDK resulted in conversion of the native dimeric structure to the tetramer assembly. Conversely, the mutation of Ala134 to Glu in PaNDK leads to conversion from tetramer to dimer assembly, indicating that a single amino acid substitution at position 134 results in an alteration of the oligomeric structure of NDK. Modeling structure of HaNDK and PaNDK, based on the crystal structure of
NDK (PaNDK) forms a tetrameric assembly. The mutation of Glu134 to Ala in HaNDK resulted in conversion of the native dimeric structure to the tetramer assembly. Conversely, the mutation of Ala134 to Glu in PaNDK leads to conversion from tetramer to dimer assembly, indicating that a single amino acid substitution at position 134 results in an alteration of the oligomeric structure of NDK. Modeling structure of HaNDK and PaNDK, based on the crystal structure of  NDK, suggested sufficient repulsion by Glu134 to disrupt dimer-dimer interaction to form tetramer.
NDK, suggested sufficient repulsion by Glu134 to disrupt dimer-dimer interaction to form tetramer.
Honjo, Eijiro; Tamada, Taro; Maeda, Yoshitake*; Koshiba, Takumi*; Matsukura, Yasuko*; Okamoto, Tomoyuki*; Ishibashi, Matsujiro*; Tokunaga, Masao*; Kuroki, Ryota
Acta Crystallographica Section F, 61(8), p.788 - 790, 2005/08
Times Cited Count:7 Percentile:54.65(Biochemical Research Methods)Granulocyte colony-stimulating factor (GCSF) receptor receives signals for regulating the proliferation and differentiation of the precursor cells of granulocytes. The complex composed of two GCSFs and two GCSF receptors was crystallized. The crystal of the complex was grown in 1.0 M sodium formate and 0.1 M sodium acetate (pH4.6). It belongs to the space group 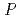 4
4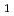 2
2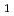 2 (or its enantiomorph
2 (or its enantiomorph 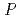 4
4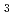 2
2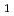 2) with unit cell dimensions of
2) with unit cell dimensions of 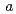 =
= 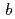 = 110.1
= 110.1  ,
, 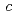 = 331.8
= 331.8  . However, the diffraction data from the crystal beyond 5
. However, the diffraction data from the crystal beyond 5  resolution could not be collected. Since the heterogeneity of GCSF receptor seems to interrupt growth of good quality crystals, the GCSF receptor was fractionated by achromatography. Crystals of GCSF/fractionated GCSF receptor complex were grown as a new crystal form in 0.2 M ammonium phosphate. The new crystal diffracts beyond 3.0
resolution could not be collected. Since the heterogeneity of GCSF receptor seems to interrupt growth of good quality crystals, the GCSF receptor was fractionated by achromatography. Crystals of GCSF/fractionated GCSF receptor complex were grown as a new crystal form in 0.2 M ammonium phosphate. The new crystal diffracts beyond 3.0  resolution and belongs to space group
resolution and belongs to space group 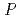 3
3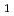 21 (or its enantiomorph
21 (or its enantiomorph 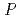 3
3 21) with unit-cell parameters
21) with unit-cell parameters 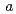 =
= 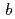 = 134.8,
= 134.8, 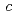 = 105.7
= 105.7  .
.
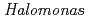 sp.593 (HaNDK)
sp.593 (HaNDK)Arai, Shigeki; Yonezawa, Yasushi; Okazaki, Nobuo; Tamada, Taro; Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Tokunaga, Masao*; Kuroki, Ryota
no journal, ,
The nucleoside diphosphate kinases (NDKs) are known to have a tetrameric or hexameric oligomer structure formed by association of common dimeric components. In order to understand the oligomeric structure of NDK, the wild-type NDK from Halomonas sp. 593 (HaNDK) was determined by X-ray crystallography. The wild-type HaNDK only exhibited a dimeric form that is the same as a common dimer unit seen in other NDKs. This is the first evidence of a dimeric structure for HaNDK. To explore the effect of mutation on the assembly form of HaNDK, E134, which is a key interface residue for other tetrameric NDKs, was replaced with Ala, and the structure was determined by X-ray crystallography. Two kinds of tetrameric assemblies, Type I and Type II, appeared in the asymmetric unit of the E134A mutant HaNDK. The change from dimeric to tetrameric assembly is attributed to the removal of negative charge repulsion caused by the E134 in the wild-type HaNDK.
 -Lactamase derived from
-Lactamase derived from 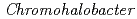 sp.560
sp.560Arai, Shigeki; Tokunaga, Hiroko*; Tamada, Taro; Yonezawa, Yasushi; Adachi, Motoyasu; Yamada, Mitsugu; Ishibashi, Matsujiro*; Tokunaga, Masao*; Kuroki, Ryota
no journal, ,
Halophilic proteins can bind various inorganic ions with its negative charges located over the molecular surface. We are investigating the molecular structure of the halophilic proteins because halophilic proteins might be used as a material capturing for the rare metals and/or the radioactive metal ions. Recently, we succeeded in X-ray crystallographic analysis of the halophilic  -Lactamase (HaBLA) derived from
-Lactamase (HaBLA) derived from 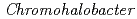 sp.560 at 3.0
sp.560 at 3.0  resolution. From this structural analysis, we found that the back bone structure and the positively charged active site feature of HaBLA was similar to those of non-halophilic BLA. The molecular surface of HaBLA were, however, occupied by large number of negative charges. This structural information is very useful to improve the specificity of metals such as Cs or Sr.
resolution. From this structural analysis, we found that the back bone structure and the positively charged active site feature of HaBLA was similar to those of non-halophilic BLA. The molecular surface of HaBLA were, however, occupied by large number of negative charges. This structural information is very useful to improve the specificity of metals such as Cs or Sr.
Adachi, Motoyasu; Arai, Shigeki; Matsumoto, Fumiko; Kuroki, Ryota; Hatanaka, Takaaki*; Ito, Yuji*; Hidaka, Koshi*; Tsuda, Yuko*; Kiso, Yoshiaki*
no journal, ,
HIV protease is known as a drug target protein. To compare interactions between HIVPR and inhibitors, single-chained HIVPR linked by a disulfide bond was designed as N98C mutant. Both WT N98C and A17 N98C mutants were expressed as inclusion body and refolded by dilution method. The yield of the purified enzymes was similar to that of WT. We will report the results of interactions between inhibitors and WT N98C and A17 N98C mutants by physicochemical and crystal structure analyses.
Adachi, Motoyasu; Hatanaka, Takaaki*; Ito, Yuji*; Hidaka, Koshi*; Tsuda, Yuko*; Kiso, Yoshiaki*; Kuroki, Ryota
no journal, ,
HIV protease is known as a drug target protein. It is important to clear relationship between structural data and kinetic and physicochemical parameters for drug design. We prepared single-chained enzyme linked by two amino acids and cross-liked enzyme bridged by disulfide bond. The two single-chained enzymes were expressed as inclusion body and refolded by dilution method. The purified enzyme complexed with inhibitor KNI-272 was crystallized, and solved the crystal structures. The designed sc- and cl-HIV-PR will be useful for evaluate the affinity of newly designed inhibitors from kinetic and thermodynamic point of view. Finally, we also report the results of analysis in affinity of inhibitors by surface plasmon resonance.
 and Sr
and Sr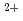 recognition mechanism of halophilic
recognition mechanism of halophilic  -lactamase
-lactamaseArai, Shigeki; Yonezawa, Yasushi; Adachi, Motoyasu; Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Tokunaga, Masao*; Kuroki, Ryota
no journal, ,
Proteins can distinguish various metal ions. Recently, we succeeded in X-ray crystallographic analysis of the halophilic  -Lactamase (HaBLA) derived from
-Lactamase (HaBLA) derived from 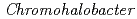 sp.560 under the existence of Cs+ and Sr2+. Diffraction data at 2.0
sp.560 under the existence of Cs+ and Sr2+. Diffraction data at 2.0  resolution (space group P31, Unit cell a = b =115.9
resolution (space group P31, Unit cell a = b =115.9  , c =67.9
, c =67.9  , Rmerge 9.6%) was collected. Three Cs
, Rmerge 9.6%) was collected. Three Cs and six Sr
and six Sr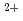 metal ion binding sites of HaBLA molecules in the asymmetric unit were identified. This structural information is very useful to create the artificial Cs
metal ion binding sites of HaBLA molecules in the asymmetric unit were identified. This structural information is very useful to create the artificial Cs or Sr
or Sr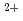 binding site on the protein molecules.
binding site on the protein molecules.
Matsumoto, Fumiko; Hatanaka, Takaaki*; Adachi, Motoyasu; Shimizu, Rumi; Tamada, Taro; Ito, Yuji*; Kuroki, Ryota
no journal, ,
no abstracts in English
Komazaki, Shinichi*; Chida, Shinji*; Nakata, Toshiya; Kono, Yutaka*; Tanigawa, Hiroyasu
no journal, ,
no abstracts in English
 -Lactamase derived from
-Lactamase derived from 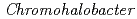 sp.560
sp.560Arai, Shigeki; Yonezawa, Yasushi; Okazaki, Nobuo; Tamada, Taro; Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Tokunaga, Masao*; Kuroki, Ryota
no journal, ,
We determined the crystal structure of halophilic  -Lactamase obtained from
-Lactamase obtained from 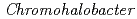 sp.560 by X-ray crystallography. Since halophilic enzymes can bind many metal ions on its molecular surface, we are tring to find the Na
sp.560 by X-ray crystallography. Since halophilic enzymes can bind many metal ions on its molecular surface, we are tring to find the Na , Mg
, Mg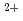 and Cs
and Cs binding sites of halophilic
binding sites of halophilic  -Lactamase from determined crystal structure.
-Lactamase from determined crystal structure.
Arai, Shigeki; Yonezawa, Yasushi*; Tamada, Taro; Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Tokunaga, Masao*; Kuroki, Ryota
no journal, ,
no abstracts in English
Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Arisaka, Fimio*; Arai, Shigeki; Kuroki, Ryota; Yamaguchi, Rui*; Arakawa, Tsutomu*; Tokunaga, Masao*
no journal, ,
no abstracts in English
Arai, Shigeki; Yonezawa, Yasushi; Okazaki, Nobuo; Tamada, Taro; Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Tokunaga, Masao*; Kuroki, Ryota
no journal, ,
The nucleoside diphosphate kinases (NDKs) are known to have a tetrameric or hexameric oligomer structure formed by association of common dimeric components. We determined the crystal structure of E134A mutant NDK from Halomonas sp. 593 (HaNDK) and found that two kinds of tetrameric assemblies, Type I seen in the Myxococcus NDK tetramer and Type II seen in the E.coli NDK tetramer, appeared in the asymmetric unit. Change in the assembly form may be an effective way for NDK to acquire molecular characteristics suited to various circumstances.
Arai, Shigeki; Yonezawa, Yasushi; Okazaki, Nobuo; Tamada, Taro; Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Tokunaga, Masao*; Kuroki, Ryota
no journal, ,
The nucleoside diphosphate kinases (NDKs) are known to have a tetrameric or hexameric oligomer structure formed by association of common dimeric components. We determined the crystal structure of E134A mutant NDK from 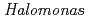 sp. 593 (HaNDK) and found that two kinds of tetrameric assemblies, Type I seen in the
sp. 593 (HaNDK) and found that two kinds of tetrameric assemblies, Type I seen in the 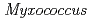 NDK tetramer and Type II seen in the
NDK tetramer and Type II seen in the 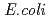 NDK tetramer, appeared in the asymmetric unit. Change in the assembly form may be an effective way for NDK to acquire molecular characteristics suited to various circumstances.
NDK tetramer, appeared in the asymmetric unit. Change in the assembly form may be an effective way for NDK to acquire molecular characteristics suited to various circumstances.