Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Nakagawa, Hiroshi; Saio, Tomohide*; Nagao, Michihiro*; Inoue, Rintaro*; Sugiyama, Masaaki*; Ajito, Satoshi; Tominaga, Taiki*; Kawakita, Yukinobu
Biophysical Journal, 120(16), p.3341 - 3354, 2021/08
Times Cited Count:4 Percentile:36.25(Biophysics)A multi-domain protein can have various conformations in solution. Interactions with other molecules result in the stabilization of one of the conformations and change in the domain dynamics. SAXS, a well-established experimental technique, can be employed to elucidate the conformation of a multi-domain protein in solution. NSE spectroscopy is a promising technique for recording the domain dynamics in nanosecond and nanometer scale. Despite the great efforts, there are still under development. Thus, we quantitatively removed the contribution of diffusion dynamics and hydrodynamic interactions from the NSE data via incoherent scattering, revealing the differences in the domain dynamics of the three functional states of a multi-domain protein, MurD. The differences among the three states can be explained by two domain modes.
Takemoto, Noriyuki; Izumo, Hironobu; Inoue, Shuichi; Abe, Shinichi; Naka, Michihiro; Akashi, Kazutomo; Omi, Masao; Miyazawa, Masataka; Baba, Osamu*; Nagao, Yoshiharu
JAEA-Review 2008-051, 36 Pages, 2008/10
The JMTR has been refurbished to restart operation in FY2011. The restarted JMTR plays roles of (1) measures for long-term operation of light water reactors, (2) improvement in scientific technique, (3) increase of industrial use, (4) training of human resources, etc. It is needed to operate the restarted JMTR safety and stably and maintain high available factor (50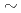 70%) because of increasing of irradiation utilization demand. In this report, measures for training of reactor operators, organization for operating, etc were proposed to operate reactor safety and smoothly. And also reactor operation procedure was examined to improve available factor up to world level for materials testing reactor. As a result, it was turned out to be possible to realize stably 210 days operation per year (available factor: 60%).
70%) because of increasing of irradiation utilization demand. In this report, measures for training of reactor operators, organization for operating, etc were proposed to operate reactor safety and smoothly. And also reactor operation procedure was examined to improve available factor up to world level for materials testing reactor. As a result, it was turned out to be possible to realize stably 210 days operation per year (available factor: 60%).
Takahashi, Nobuaki; Nishida, Koji*; Inoue, Rintaro*; Ogawa, Hiroki*; Kanaya, Toshiji*; Nagao, Michihiro*
NSL News Letter, 2007-4, p.155 - 157, 2007/04
We have studied dynamics of poly(vinyl alcohol) (PVA) gel in a mixture of deuterated dimethyl sulfoxide (DMSO-d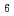 ) and D
) and D O (60/40 by volume) during heating process from 25
O (60/40 by volume) during heating process from 25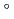 C to 80
C to 80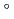 C using neutron spin-echo (NSE) techniques.
C using neutron spin-echo (NSE) techniques.
Takahashi, Nobuaki; Nishida, Koji*; Tsubouchi, Tsuyoshi*; Ogawa, Hiroki*; Inoue, Rintaro*; Kanaya, Toshiji*; Nagao, Michihiro*
ISSP Activity Report on Neutron scattering Research; Experimental Reports (CD-ROM), 13, 2 Pages, 2006/00
no abstracts in English
Kanaya, Toshiji; Takahashi, Nobuaki; Inoue, Rintaro*; Matsuba, Go*; Nishida, Koji*; Nagao, Michihiro*
ISSP Activity Report on Neutron scattering Research; Experimental Reports (CD-ROM), 13, 1 Pages, 2006/00
Ouchi, Nobuo; Mizumoto, Motoharu; Kusano, Joichi; Chishiro, Etsuji; Hasegawa, Kazuo; Akaoka, Nobuo*; Saito, Kenji*; Noguchi, Shuichi*; Kako, Eiji*; Inoue, Hitoshi*; et al.
Proceedings of 20th International Linac Conference (CD-ROM), 1 Pages, 2000/00
no abstracts in English
Nakagawa, Hiroshi; Saio, Tomohide*; Sugiyama, Masaaki*; Inoue, Rintaro*; Nagao, Michihiro*
no journal, ,
It is necessary for the understanding of structure polymorphism of various molecules and interacting protein and the plastic molecules base to clarify the fluctuation of the domain as the mer. In this study, I targeted protein MurD consisting of three domains and a ligand was in a free state and analyzed domain exercise by the quantum beam dispersion method using X-rays and the neutron and the correlation structure analytical method that fused of the molecules simulation about an ATP-binding state, three states of the Compound1-binding state. By the solution small angle scattering experiment, I showed that the solution structure of the 3 state was different. In addition, I showed good agreement with the dispersion profile obtained from the molecular simulation and confirmed that I could discuss solution structure in atom resolving power from experimental data of the low resolving power. I extracted a domain campaign in conjunction with the function from the chief ingredient analysis of the molecular simulation result. Furthermore, a result to suggest what a fluctuation and a couple of the amino acid residue that this domain campaign participated in the combination with interaction molecules of MurD did was provided. By the method that fused by an experiment method of dynamic solution structure analysis including a neutron spin echo and technique of the calculation science in addition to small angle scattering, I visualize domain exercise of protein in atom resolving power and, in the announcement, argue by a function of the protein from the dynamic structure that a couple did between the hierarchies of different space scale.
Nakagawa, Hiroshi; Saio, Tomohide*; Nagao, Michihiro*; Inoue, Rintaro*; Sugiyama, Masaaki*; Tominaga, Taiki*; Kawakita, Yukinobu
no journal, ,
The flexible conformation of a multidomain protein is responsible for its biological function. 3-domain protein: MurD (47kDa) changes its domain conformation sequentially from open to semi-closed to closed conformation during enzymatic reactions. However, the dynamics of the domains in each conformation is unknown. In this study, we combined small-angle X-ray and neutron scattering (SAXS and SANS), dynamic light scattering (DLS), neutron back scattering (NBS), neutron spin echo (NSE), and molecular dynamics (MD) simulations to investigate the conformational dynamics of MurD in the three corresponding states (apo and ATP, inhibitor-bound state). The analysis showed that the conformational dynamics of MurD during the enzymatic reaction was not affected. The analysis suggests that the changes in domain dynamics during enzymatic reactions are related to the affinity and reaction efficiency with ligands that bind specifically to each reaction state.
Nakagawa, Hiroshi; Saio, Tomohide*; Nagao, Michihiro*; Inoue, Rintaro*; Sugiyama, Masaaki*; Tominaga, Taiki*; Kawakita, Yukinobu
no journal, ,
The flexible conformation of a multidomain protein is responsible for its biological function. 3-domain protein: MurD (47kDa) changes its domain conformation sequentially from open to semi-closed to closed conformation during enzymatic reactions. However, the dynamics of the domains in each conformation is unknown. In this study, we combined small-angle X-ray and neutron scattering (SAXS and SANS), dynamic light scattering (DLS), neutron back scattering (NBS), neutron spin echo (NSE), and molecular dynamics (MD) simulations to investigate the conformational dynamics of MurD in the three corresponding states (apo and ATP, inhibitor-bound state). The analysis showed that the conformational dynamics of MurD during the enzymatic reaction was not affected. The analysis suggests that the changes in domain dynamics during enzymatic reactions are related to the affinity and reaction efficiency with ligands that bind specifically to each reaction state.
Asai, Takafumi*; Inoue, Chihiro*; Toyonaga, Keita*; Jinno, Satoshi; Ryazantsev, S.*; Pikuz, T.*; Yamauchi, Tomoya*; Kanasaki, Masato*; Fukuda, Yuji*
no journal, ,
The J-KAREN-P laser at the Kansai Photon Science Institute was irradiated to a cluster target and confirmed the second harmonic generation. Polarization measurements of the generated second harmonic revealed that the polarization plane of the component that passed through the laser-generated plasma was rotated. This result suggests that a vortex-like magnetic field centered on the laser axis is generated.