Predicting cetuximab accumulation in 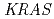 wild-type and
wild-type and 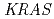 mutant colorectal cancer using
mutant colorectal cancer using 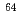 Cu-labeled cetuximab positron emission tomography
Cu-labeled cetuximab positron emission tomography
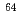 Cu標識セツキシマブPETによるKRAS野生型及びKRAS変異型大腸癌へのセツキシマブ集積の予測
Cu標識セツキシマブPETによるKRAS野生型及びKRAS変異型大腸癌へのセツキシマブ集積の予測
Achmad, A.*; 花岡 宏史*; 吉岡 弘樹*; 山元 進司*; 富永 英之*; 荒木 拓也*; 大島 康宏; 織内 昇*; 遠藤 啓吾*
Achmad, A.*; Hanaoka, Hirofumi*; Yoshioka, Hiroki*; Yamamoto, Shinji*; Tominaga, Hideyuki*; Araki, Takuya*; Ohshima, Yasuhiro; Oriuchi, Noboru*; Endo, Keigo*
Overexpression of epidermal growth factor receptor (EGFR) is common in colorectal cancer. However, cetuximab, an EGFR-targeting drug, is useful only for a subset of patients and no single predictor other than V-Ki-ras2 Kirsten rat sarcoma viral oncogene homolog (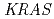 ) mutation status has been established. In this study, we investigated cetuximab accumulation in colorectal tumors using
) mutation status has been established. In this study, we investigated cetuximab accumulation in colorectal tumors using 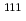 In-DOTA-cetuximab, and evaluated the potential of positron emission tomography (PET) imaging of
In-DOTA-cetuximab, and evaluated the potential of positron emission tomography (PET) imaging of 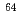 Cu-DOTA-cetuximab. We found that
Cu-DOTA-cetuximab. We found that 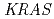 wild-type tumors had significantly higher
wild-type tumors had significantly higher  In-DOTA-cetuximab accumulation than
In-DOTA-cetuximab accumulation than 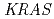 mutant tumors. Based on
mutant tumors. Based on 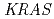 mutation status, a strong correlation was found between
mutation status, a strong correlation was found between  In-DOTA-cetuximab tumor uptake and EGFR expression level. Significant correlation was also found between tumor uptake of
In-DOTA-cetuximab tumor uptake and EGFR expression level. Significant correlation was also found between tumor uptake of 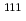 In-DOTA-cetuximab and
In-DOTA-cetuximab and 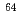 Cu-DOTA-cetuximab. PET imaging with
Cu-DOTA-cetuximab. PET imaging with 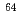 Cu-DOTA-cetuximab effectively visualized cetuximab accumulation in colorectal tumors with a wide variety of EGFR expression levels and different
Cu-DOTA-cetuximab effectively visualized cetuximab accumulation in colorectal tumors with a wide variety of EGFR expression levels and different 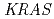 mutation status as commonly encountered in the clinical setting. Our findings suggest that this radioimmunoimaging can be clinically translated as an in vivo tool to predict cetuximab accumulation in colorectal cancer patients prior to cetuximab therapy.
mutation status as commonly encountered in the clinical setting. Our findings suggest that this radioimmunoimaging can be clinically translated as an in vivo tool to predict cetuximab accumulation in colorectal cancer patients prior to cetuximab therapy.