Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
Initialising ...
 binding site of a
binding site of a  -lactamase from the halophile by anomalous X-ray diffraction
-lactamase from the halophile by anomalous X-ray diffractionArai, Shigeki; Shibazaki, Chie; Shimizu, Rumi; Adachi, Motoyasu; Tamada, Taro; Tokunaga, Hiroko*; Ishibashi, Matsujiro*; Tokunaga, Masao*; Kuroki, Ryota
Kyushu Shinkurotoronko Kenkyu Senta Nempo, 2014, p.17 - 19, 2016/03
no abstracts in English
 sp. ME121, isolated from soil as a mixed single colony with
sp. ME121, isolated from soil as a mixed single colony with  sp. 32K
sp. 32KFujinami, Shun*; Takeda, Kiyoko*; Onodera, Takefumi*; Sato, Katsuya; Shimizu, Tetsu*; Wakabayashi, Yu*; Narumi, Issey*; Nakamura, Akira*; Ito, Masahiro*
Genome Announcements (Internet), 3(5), p.e01005-15_1 - e01005-15_2, 2015/09
 -lactamase from the moderate halophile
-lactamase from the moderate halophile  sp.560 and the discovery of a Cs
sp.560 and the discovery of a Cs -selective binding site
-selective binding siteArai, Shigeki; Yonezawa, Yasushi*; Okazaki, Nobuo*; Matsumoto, Fumiko*; Shibazaki, Chie; Shimizu, Rumi; Yamada, Mitsugu*; Adachi, Motoyasu; Tamada, Taro; Kawamoto, Masahide*; et al.
Acta Crystallographica Section D, 71(3), p.541 - 554, 2015/03
Times Cited Count:8 Percentile:52.42(Biochemical Research Methods)The crystal structure of halophilic  -lactamase from
-lactamase from  sp.560 (HaBLA) was determined using X-ray crystallography. Moreover, the locations of bound Sr
sp.560 (HaBLA) was determined using X-ray crystallography. Moreover, the locations of bound Sr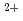 and Cs
and Cs ions were identified by anomalous X-ray diffraction. The location of one Cs
ions were identified by anomalous X-ray diffraction. The location of one Cs specific binding site was identified on HaBLA even in the presence of 9-fold molar excess of Na
specific binding site was identified on HaBLA even in the presence of 9-fold molar excess of Na (90 mM Na
(90 mM Na /10 mM Cs
/10 mM Cs ). This Cs
). This Cs binding site is formed by two main-chain O atoms and an aromatic ring of a side chain of Trp. An aromatic ring of Trp interacts with Cs
binding site is formed by two main-chain O atoms and an aromatic ring of a side chain of Trp. An aromatic ring of Trp interacts with Cs by the cation-
by the cation-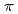 interaction. The observation of a selective and high-affinity Cs
interaction. The observation of a selective and high-affinity Cs binding site provides important information that is useful for designing artificial Cs
binding site provides important information that is useful for designing artificial Cs binding sites useful in bioremediation of radioactive isotopes.
binding sites useful in bioremediation of radioactive isotopes.

Adachi, Motoyasu; Hirayama, Hiroshi; Shimizu, Rumi; Sato, Katsuya; Narumi, Issey*; Kuroki, Ryota
Protein Science, 23(10), p.1349 - 1358, 2014/10
Times Cited Count:10 Percentile:26.13(Biochemistry & Molecular Biology)Pleiotropic protein promoting DNA repair A (PprA) is a key protein that facilitates the extreme radioresistance of  . To clarify the role of PprA in the radioresistance mechanism, the interaction between recombinant PprA expressed in Escherichia coli with several double-stranded DNAs was investigated. In a gel-shift assay, the band shift of supercoiled pUC19 DNA caused by the binding of PprA showed a bimodal distribution, which was promoted by the addition of 1 mM Mg, Ca, or Sr ions. The dissociation constant of the PprA-supercoiled pUC19 DNA complex, calculated from the relative portions of shifted bands, was 0.6
. To clarify the role of PprA in the radioresistance mechanism, the interaction between recombinant PprA expressed in Escherichia coli with several double-stranded DNAs was investigated. In a gel-shift assay, the band shift of supercoiled pUC19 DNA caused by the binding of PprA showed a bimodal distribution, which was promoted by the addition of 1 mM Mg, Ca, or Sr ions. The dissociation constant of the PprA-supercoiled pUC19 DNA complex, calculated from the relative portions of shifted bands, was 0.6 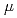 M with a Hill coefficient of 3.3 in the presence of 1 mM Mg acetate. This indicates that at least 281 PprA molecules are required to saturate a supercoiled pUC19 DNA, which is consistent with the number of bound PprA molecules estimated by the UV absorption of the PprA-pUC19 complex purified by gel filtration. This saturation also suggests linear polymerization of PprA along the dsDNA. On the other hand, the bands of linear dsDNA and nicked circular dsDNA that eventually formed PprA complexes did not saturate, but created larger molecular complexes when the PprA concentration was greater than 1.3
M with a Hill coefficient of 3.3 in the presence of 1 mM Mg acetate. This indicates that at least 281 PprA molecules are required to saturate a supercoiled pUC19 DNA, which is consistent with the number of bound PprA molecules estimated by the UV absorption of the PprA-pUC19 complex purified by gel filtration. This saturation also suggests linear polymerization of PprA along the dsDNA. On the other hand, the bands of linear dsDNA and nicked circular dsDNA that eventually formed PprA complexes did not saturate, but created larger molecular complexes when the PprA concentration was greater than 1.3 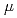 M. This result implies that DNA-bound PprA aids association of the termini of damaged DNAs, which is regulated by the concentration of PprA.
M. This result implies that DNA-bound PprA aids association of the termini of damaged DNAs, which is regulated by the concentration of PprA.
Ninomiya, Kazuaki*; Omote, Sayuri*; Sato, Katsuya; Narumi, Issey*; Shimizu, Nobuaki*
JAEA-Review 2013-059, JAEA Takasaki Annual Report 2012, P. 115, 2014/03
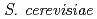
Matsuo, Yoichiro*; Izumi, Yoshinobu*; Hase, Yoshihiro; Sakamoto, Ayako; Nozawa, Shigeki; Narumi, Issey*; Shimizu, Kikuo*
JAEA-Review 2013-059, JAEA Takasaki Annual Report 2012, P. 112, 2014/03
Adachi, Motoyasu; Shimizu, Rumi; Kuroki, Ryota; Blaber, M.
Journal of Synchrotron Radiation, 20(6), p.953 - 957, 2013/11
Times Cited Count:2 Percentile:12.49(Instruments & Instrumentation)Symfoil-4P is a 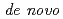 protein exhibiting the threefold symmetrical beta-trefoil fold designed based on the human acidic fibroblast growth factor. First three asparagine-glycine sequences of Symfoil-4P are replaced with glutamine-glycine (Symfoil-QG) or serine-glycine (Symfoil-SG) sequences protecting from deamidation, and His-Symfoil-II was prepared by introducing a protease digestion site into Symfoil-QG so that Symfoil-II has three complete repeats after removal of the N-terminal histidine tag. The Symfoil-QG and SG and His-Symfoil-II proteins were expressed in
protein exhibiting the threefold symmetrical beta-trefoil fold designed based on the human acidic fibroblast growth factor. First three asparagine-glycine sequences of Symfoil-4P are replaced with glutamine-glycine (Symfoil-QG) or serine-glycine (Symfoil-SG) sequences protecting from deamidation, and His-Symfoil-II was prepared by introducing a protease digestion site into Symfoil-QG so that Symfoil-II has three complete repeats after removal of the N-terminal histidine tag. The Symfoil-QG and SG and His-Symfoil-II proteins were expressed in 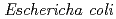 as soluble protein, and purified by nickel affinity chromatography. Symfoil-II was further purified by anion-exchange chromatography after removing the HisTag by proteolysis. Symfoil-QG and II crystals gave 1.5 and 1.1
as soluble protein, and purified by nickel affinity chromatography. Symfoil-II was further purified by anion-exchange chromatography after removing the HisTag by proteolysis. Symfoil-QG and II crystals gave 1.5 and 1.1 , resolution, respectively. The refined crystal structure of Symfoil-II showed pseudo-threefold symmetry as expected from other Symfoils.
, resolution, respectively. The refined crystal structure of Symfoil-II showed pseudo-threefold symmetry as expected from other Symfoils.
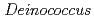 spp. under the ISS environmental conditions for performing exposure experiments of microbes in the Tanpopo mission
spp. under the ISS environmental conditions for performing exposure experiments of microbes in the Tanpopo missionKawaguchi, Yuko*; Yang, Y.*; Kawashiri, Narutoshi*; Shiraishi, Keisuke*; Takasu, Masako*; Narumi, Issey*; Sato, Katsuya; Hashimoto, Hirofumi*; Nakagawa, Kazumichi*; Tanigawa, Yoshiaki*; et al.
Origins of Life and Evolution of Biospheres, 43(4-5), p.411 - 428, 2013/10
Times Cited Count:42 Percentile:79.35(Biology)Matsuo, Yoichiro*; Izumi, Yoshinobu*; Hase, Yoshihiro; Sakamoto, Ayako; Nozawa, Shigeki; Narumi, Issei; Shimizu, Kikuo*
JAEA-Review 2012-046, JAEA Takasaki Annual Report 2011, P. 105, 2013/01
Ninomiya, Kazuaki*; Nomura, Tomoyo*; Sato, Katsuya; Narumi, Issei; Shimizu, Nobuaki*
JAEA-Review 2012-046, JAEA Takasaki Annual Report 2011, P. 108, 2013/01
Ninomiya, Kazuaki*; Soda, Hiroshi*; Sato, Katsuya; Narumi, Issei; Shimizu, Nobuaki*
JAEA-Review 2011-043, JAEA Takasaki Annual Report 2010, P. 111, 2012/01
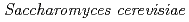
Matsuo, Yoichiro*; Izumi, Yoshinobu*; Hase, Yoshihiro; Sakamoto, Ayako; Nozawa, Shigeki; Narumi, Issei; Shimizu, Kikuo*
JAEA-Review 2011-043, JAEA Takasaki Annual Report 2010, P. 108, 2012/01
Ninomiya, Kazuaki*; Soda, Hiroshi*; Sato, Katsuya; Narumi, Issei; Shimizu, Nobuaki*
JAEA-Review 2010-065, JAEA Takasaki Annual Report 2009, P. 78, 2011/01
no abstracts in English
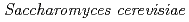 cells
cellsShimizu, Kikuo*; Matsuo, Yoichiro*; Izumi, Yoshinobu*; Hase, Yoshihiro; Nozawa, Shigeki; Sakamoto, Ayako; Narumi, Issei
JAEA-Review 2010-065, JAEA Takasaki Annual Report 2009, P. 79, 2011/01
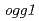 and
and 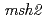 mutants
mutantsMatsuo, Yoichiro*; Nishijima, Shigehiro*; Hase, Yoshihiro; Nozawa, Shigeki; Sakamoto, Ayako; Narumi, Issei; Shimizu, Kikuo*
JAEA-Review 2009-041, JAEA Takasaki Annual Report 2008, P. 75, 2009/12
no abstracts in English
Okamura, Masachika*; Shimizu, Akira*; Onishi, Noboru*; Hase, Yoshihiro; Yoshihara, Ryohei; Narumi, Issei
JAEA-Review 2008-055, JAEA Takasaki Annual Report 2007, P. 62, 2008/11
no abstracts in English
Matsuo, Yoichiro*; Nishijima, Shigehiro*; Hase, Yoshihiro; Sakamoto, Ayako; Narumi, Issei; Shimizu, Kikuo*
JAEA-Review 2008-055, JAEA Takasaki Annual Report 2007, P. 60, 2008/11
no abstracts in English
 by heavy ion irradiation
by heavy ion irradiationSakai, Kazuya*; Kobayashi, Hiroyuki*; Shimizu, Masatoshi*; Oshima, Akio*; Kato, Shigeaki*; Sato, Katsuya; Narumi, Issei
JAEA-Review 2008-055, JAEA Takasaki Annual Report 2007, P. 85, 2008/11
no abstracts in English
Takahashi, Shinya*; Nakagawa, Mayu; Tanaka, Atsushi; Narumi, Issei; Shimizu, Kikuo*; Sakamoto, Ayako
JAEA-Review 2008-055, JAEA Takasaki Annual Report 2007, P. 58, 2008/11
no abstracts in English
Matsuo, Yoichiro*; Nishijima, Shigehiro*; Hase, Yoshihiro; Sakamoto, Ayako; Yokota, Yuichiro; Narumi, Issei; Shimizu, Kikuo*
JAEA-Review 2007-060, JAEA Takasaki Annual Report 2006, P. 86, 2008/03
no abstracts in English